---
title: "Coronavirus in Kenya"
author: "University of Nairobi"
output:
flexdashboard::flex_dashboard:
orientation: rows
source_code: embed
vertical_layout: fill
self_contained: FALSE
---
```{r setup, include=FALSE}
#------------------ Packages ------------------
knitr::opts_chunk$set(echo = FALSE)
library(tidyverse)
library(EpiEstim)
library(incidence)
library(deSolve)
library(lubridate)
library(plotly)
library(dplyr)
library(reshape)
library(htmltools)
library(magrittr)
library(stringr)
library(tibble)
library(readxl)
library(lubridate)
library(rgdal)
library(sf)
library(gghighlight)
library(magrittr)
library(plotly)
library(janitor)
#library(broom)
#library(hrbrthemes)
library(rvest)
library(gt)
library(deSolve)
library(distcrete)
library(epitrix)
library(projections)
library(earlyR)
library(flexdashboard)
#library(squire)
#install.packages("devtools")
# install.packages("devtools")
#devtools::install_github("RamiKrispin/coronavirus")
#devtools::install_github("RamiKrispin/coronavirus", force = TRUE)
#install.packages("coronavirus")
library(coronavirus)
data(coronavirus)
#coronavirus<-refresh_coronavirus_jhu()
#update_dataset()
#View(coronavirus)
#max(coronavirus$date)
`%>%` <- magrittr::`%>%`
#------------------ Parameters ------------------
# Set colors
# https://www.w3.org/TR/css-color-3/#svg-color
confirmed_color <- "black"
active_color <- "#1f77b4"
recovered_color <- "forestgreen"
death_color <- "red"
#------------------ Data ------------------
df <- coronavirus %>%
dplyr::filter(date >= "2020-03-12") %>%
dplyr::filter(country == "Kenya") %>%
dplyr::group_by(country, type) %>%
dplyr::summarise(total = sum(cases)) %>%
tidyr::pivot_wider(
names_from = type,
values_from = total
) %>%
# dplyr::mutate(unrecovered = confirmed - ifelse(is.na(recovered), 0, recovered) - ifelse(is.na(death), 0, death)) %>%
dplyr::mutate(unrecovered = confirmed - ifelse(is.na(death), 0, death)) %>%
dplyr::arrange(-confirmed) %>%
dplyr::ungroup() %>%
dplyr::mutate(country = dplyr::if_else(country == "United Arab Emirates", "UAE", country)) %>%
dplyr::mutate(country = dplyr::if_else(country == "Mainland China", "China", country)) %>%
dplyr::mutate(country = dplyr::if_else(country == "North Macedonia", "N.Macedonia", country)) %>%
dplyr::mutate(country = trimws(country)) %>%
dplyr::mutate(country = factor(country, levels = country))
#worlddatacovid <- read_csv("https://covid.ourworldindata.org/data/owid-covid-data.csv") # data from ourworlddata
#ecdccovid <- read_csv("https://opendata.ecdc.europa.eu/covid19/casedistribution/csv", na.strings = "", fileEncoding = "UTF-8-BOM")
#humdatacovid <- read_csv("https://datahub.io/core/covid-19/r/time-series-19-covid-combined.csv")
# df_daily <- humdatacovid %>%
# dplyr::filter(Date >= "2020-03-12") %>%
# dplyr::filter(`Country/Region` == "Kenya") %>%
# #dplyr::group_by(Date) %>%
# dplyr::mutate(
# confirmed_cum = cumsum(Confirmed),
# recovered_cum = cumsum(Recovered),
# death_cum = cumsum(Deaths),
# active = (confirmed_cum - death_cum - recovered_cum),
# active_cum = cumsum(active)) %>%
# dplyr::ungroup()
df_daily <- coronavirus %>%
dplyr::filter(date >= "2020-03-10") %>%
dplyr::filter(country == "Kenya") %>%
dplyr::group_by(date, type) %>%
dplyr::summarise(total = sum(cases, na.rm = TRUE)) %>%
tidyr::pivot_wider(
names_from = type,
values_from = total
) %>%
dplyr::arrange(date) %>%
dplyr::ungroup() %>%
#dplyr::mutate(active = confirmed - death - recovered) %>%
dplyr::mutate(active = confirmed - death) %>%
dplyr::mutate(recovery1=active - dplyr::lag(active))%>%
dplyr::mutate(recovery1=ifelse(is.na(recovery1), 0, recovery1))%>%
dplyr::mutate(
confirmed_cum = cumsum(confirmed),
recovered_cum = cumsum(recovery),
death_cum = cumsum(death),
active_cum = cumsum(active),
recovered1_cum=cumsum(abs(recovery1))
)%>%
mutate(recovered_cum=ifelse(date>as.Date("2021-08-04"), (119277+recovered1_cum), recovered_cum))
#df1 <- coronavirus %>% dplyr::filter(date == max(date))
# kecovidcases <- read_csv("kecovidincidence.csv") %>%
# dplyr::mutate(Dates = as.Date(Dates, "%d/%m/%Y")) %>%
# filter(Dates<="2020-05-13")
#
# datetoday <- today()-1
# cases <- sum(kecovidcases$Incidence)
```
# Summary {data-navmenu="Summary"}
## Row {data-width="400"}
### confirmed {.value-box}
```{r}
valueBox(
value = paste(format(sum(df$confirmed), big.mark = ","), "", sep = " "),
caption = "Total confirmed cases",
icon = "fas fa-user-md",
color = confirmed_color
)
```
### death {.value-box}
```{r}
#sum(df$death
valueBox(
value = paste(format(sum(df$death, na.rm = TRUE), big.mark = ","), " (",
round(100 * sum(df$death, na.rm = TRUE) / sum(df$confirmed), 1),
"%)",
sep = ""
),
caption = "Death cases (death rate)",
icon = "fas fa-heart-broken",
color = death_color
)
```
### Recoveries {.value-box}
```{r}
#df$recovered
valueBox(
value = paste(format(max(df_daily$recovered_cum), big.mark = ","), "", sep = " "),
caption = "Total recovered cases",
icon = "fas fa-user-md",
color = recovered_color
)
```
## Row
### **Daily cumulative cases by type for Kenya. Kindly hover on the confirmed cases for non-pharmaceutical interventions**\*
```{r}
data<- read.csv("~/Dropbox/Research and analysis/covid19/COVID19_shared/Govt_kecovidincidence.csv")%>%
select(Dates,Interventions)%>%
mutate(Dates=as.Date(Dates, "%d/%m/%Y"))%>%
mutate(Interventions=ifelse(Interventions%in%"", "None", Interventions))
df_daily<- df_daily%>%
left_join(data, by=c("date"="Dates"))%>%
mutate(Interventions=ifelse(is.na(Interventions) & date%in%as.Date("2020-03-18"), "School closure \n travel ban \n Self quarantine", ifelse(is.na(Interventions) &date%in%as.Date("2020-03-23"), "Closure of \nsocial spaces", ifelse(is.na(Interventions) &date%in%as.Date("2020-03-25"), "Nationwide \n curfew", ifelse(is.na(Interventions) & date%in%as.Date("2020-04-04"),"Wearing masks \npublic spaces", ifelse(is.na(Interventions) & date%in%as.Date("2020-04-06"), "Travel restrictions \n(Nairobi,Mombasa, \nKilifi,Kwale)",ifelse(is.na(Interventions) & date%in%as.Date("2020-04-27"), "Restaurants \nre-open", ifelse(is.na(Interventions) & date%in%as.Date("2020-05-06"), "Restricted movement & \nSuspension of public transport in \nEastleigh/Old town", ifelse(is.na(Interventions) & date%in%as.Date("2020-05-16"),"Cessation of \npersonal/public \npassenger vehicle movement \nKenya-Tanzania, \nKenya-Somalia border", ifelse(is.na(Interventions) & date%in%as.Date("2020-06-06"),"Travel restrictions \nCurfew extended", ifelse(is.na(Interventions) & date%in%as.Date("2020-07-06"),"Travel restriction lifted \nCurfew extended", ifelse(is.na(Interventions) & date%in%as.Date("2020-07-27"), "Curfew extended \n 30days", ifelse(is.na(Interventions) & date%in%as.Date("2020-08-26"),"Curfew extended \n 30days", ifelse(is.na(Interventions) & date%in%as.Date("2020-09-28"), "Bar restrictions lifted \nCurfew hours shortened",ifelse(is.na(Interventions) & date%in%as.Date("2021-03-26"),"Travel restrictions for Nairobi, Kiambu, \nMachakos, Nakuru & Kajiado", ifelse(is.na(Interventions) & date%in%as.Date("2021-06-17"), "Travel restrictions: \nBomet, Bungoma, Busia, Homa-Bay, \nKakamega, Kericho, Kisii, Kisumu, Migori, \nNyamira, Siaya, Trans-Nzoia, Vihiga", Interventions))))))))))))))))
df_daily$Interventions1<- ifelse(is.na(df_daily$Interventions)| df_daily$Interventions%in%"None", "Yes", "No")
plotly::plot_ly(data = df_daily) %>%
plotly::add_trace(
x = ~date,
# y = ~active_cum,
y = ~confirmed_cum,
text =~Interventions,
type = "scatter",
mode = "lines+markers",
# name = "Active",
name = "Confirmed",
hovertext=paste("Npi :", df_daily$Interventions,
"
Date:", df_daily$date,
"
Cumulative cases:", df_daily$confirmed_cum),
line = list(color = confirmed_color),
marker = list(color = confirmed_color)
) %>%
plotly::add_trace(
x = ~date,
# y = ~active_cum,
y = ~recovered_cum,
type = "scatter",
mode = "lines+markers",
# name = "Active",
name = "Recovered",
line = list(color = recovered_color),
marker = list(color = recovered_color)
) %>%
plotly::add_trace(
x = ~date,
y = ~death_cum,
type = "scatter",
mode = "lines+markers",
name = "Death",
line = list(color = death_color),
marker = list(color = death_color)
) %>%
plotly::layout(
title = "",
yaxis = list(title = "Cumulative number of cases"),
xaxis = list(title = "Date"),
legend = list(x = 0.1, y = 0.9),
hovermode = "compare"
)
```
```{r}
linelist<- read.csv("linelist.csv")%>%
mutate(sub_county=str_to_title(sub_county))%>%
mutate(sub_county=recode(sub_county, "Wstland"="Westlands", "West Lands"="Westlands", "Westland"="Westlands", "Zimmerman"="Roysambu", "Wote"="Makueni", "Wjr East"="Wajir East", "Wiyumiririe"="Ngobit", "Webuye"="Webuye East", "Wangige"="Kabete","Wamunyu"="Mwala", "Wamba"="Samburu East", "Wamagana"="Tetu", "Kabete"="Uthiru", "Valley"="Kilimani", "Utawala"="Embakasi East", "Upperhill"="Dagoretti North", "Umoja"="Embakasi West", "Uon"="Starehe","Uasin Gishu Town"="Eldoret North", "Uasin Gishu"="Eldoret North", "Transzoia East"="Trans Nzoia East", "Transnzoia East"="Trans Nzoia East"))%>%
mutate(sub_county=ifelse(sub_county%in%c("Yokosukacity"), NA, sub_county))
wards<- sf::st_read("gadm36_KEN_3.shp")
```
# Comparison of cases {data-navmenu="Summary"}
## Row {data-width="400"}
### **Daily new confirmed cases**
```{r}
daily_confirmed <- coronavirus %>%
dplyr::filter(type == "confirmed") %>%
dplyr::filter(date >= "2020-02-29") %>%
dplyr::mutate(country = country) %>%
dplyr::group_by(date, country) %>%
dplyr::summarise(total = sum(cases)) %>%
dplyr::ungroup() %>%
tidyr::pivot_wider(names_from = country, values_from = total)
#----------------------------------------
# Plotting the data
daily_confirmed %>%
dplyr::filter(Uganda >= 0) %>%
plotly::plot_ly() %>%
plotly::add_trace(
x = ~date,
y = ~Kenya,
type = "scatter",
mode = "lines+markers",
name = "Kenya"
) %>%
plotly::add_trace(
x = ~date,
y = ~Ethiopia,
type = "scatter",
mode = "lines+markers",
name = "Ethiopia"
) %>%
plotly::add_trace(
x = ~date,
y = ~Somalia,
type = "scatter",
mode = "lines+markers",
name = "South Sudan"
) %>%
plotly::add_trace(
x = ~date,
y = ~Tanzania,
type = "scatter",
mode = "lines+markers",
name = "Tanzania"
) %>%
plotly::add_trace(
x = ~date,
y = ~Uganda,
type = "scatter",
mode = "lines+markers",
name = "Uganda"
) %>%
plotly::layout(
title = "",
legend = list(x = 0.1, y = 0.9),
yaxis = list(title = "Number of new confirmed cases"),
xaxis = list(title = "Date"),
# paper_bgcolor = "black",
# plot_bgcolor = "black",
# font = list(color = 'white'),
hovermode = "compare",
margin = list(
# l = 60,
# r = 40,
b = 10,
t = 10,
pad = 2
)
)
```
## Row
### **Cases and deaths by country**
```{r daily_summary}
df_EU <- coronavirus %>%
# dplyr::filter(date == max(date)) %>%
dplyr::filter(country == "Kenya" |
country == "Uganda" |
country == "Tanzania" |
country == "Burundi" |
country == "South Sudan" |
country == "Ethiopia" |
country == "Rwanda") %>%
dplyr::group_by(country, type) %>%
dplyr::summarise(total = sum(cases)) %>%
tidyr::pivot_wider(
names_from = type,
values_from = total
) %>%
# dplyr::mutate(unrecovered = confirmed - ifelse(is.na(recovered), 0, recovered) - ifelse(is.na(death), 0, death)) %>%
dplyr::mutate(unrecovered = confirmed - ifelse(is.na(death), 0, death)) %>%
dplyr::arrange(confirmed) %>%
dplyr::ungroup() %>%
dplyr::mutate(country = dplyr::if_else(country == "United Arab Emirates", "UAE", country)) %>%
dplyr::mutate(country = dplyr::if_else(country == "Mainland China", "China", country)) %>%
dplyr::mutate(country = dplyr::if_else(country == "North Macedonia", "N.Macedonia", country)) %>%
dplyr::mutate(country = trimws(country)) %>%
dplyr::mutate(country = factor(country, levels = country))
plotly::plot_ly(
data = df_EU,
x = ~country,
# y = ~unrecovered,
y = ~ confirmed,
# text = ~ confirmed,
# textposition = 'auto',
type = "bar",
name = "Confirmed",
marker = list(color = active_color)
) %>%
plotly::add_trace(
y = ~death,
# text = ~ death,
# textposition = 'auto',
name = "Death",
marker = list(color = death_color)
) %>%
plotly::layout(
barmode = "stack",
yaxis = list(title = "Total cases"),
xaxis = list(title = ""),
hovermode = "compare",
margin = list(
# l = 60,
# r = 40,
b = 10,
t = 10,
pad = 2
)
)
```
# Estimated R0 {data-navmenu="Summary"}
## Row
### **Estimated R-effective**\*
```{r, echo=F, message=F, warning=F, fig.width=16, fig.height=10}
df <- coronavirus %>%
dplyr::filter(date >= "2020-03-12") %>%
dplyr::filter(country == "Kenya") %>%
dplyr::group_by(country, type) %>%
dplyr::summarise(total = sum(cases, na.rm = T)) %>%
tidyr::pivot_wider(
names_from = type,
values_from = total
) %>%
# dplyr::mutate(unrecovered = confirmed - ifelse(is.na(recovered), 0, recovered) - ifelse(is.na(death), 0, death)) %>%
dplyr::mutate(unrecovered = confirmed - ifelse(is.na(death), 0, death)) %>%
dplyr::arrange(-confirmed) %>%
dplyr::ungroup() %>%
dplyr::mutate(country = dplyr::if_else(country == "United Arab Emirates", "UAE", country)) %>%
dplyr::mutate(country = dplyr::if_else(country == "Mainland China", "China", country)) %>%
dplyr::mutate(country = dplyr::if_else(country == "North Macedonia", "N.Macedonia", country)) %>%
dplyr::mutate(country = trimws(country)) %>%
dplyr::mutate(country = factor(country, levels = country))
df_daily <- coronavirus %>%
dplyr::filter(date >= "2020-03-13") %>%
dplyr::filter(country == "Kenya") %>%
dplyr::group_by(date, type) %>%
dplyr::summarise(total = sum(cases, na.rm = TRUE)) %>%
tidyr::pivot_wider(
names_from = type,
values_from = total
) %>%
dplyr::arrange(date) %>%
dplyr::ungroup() %>%
#dplyr::mutate(active = confirmed - death - recovered) %>%
dplyr::mutate(active = confirmed - death) %>%
dplyr::mutate(
confirmed_cum = cumsum(confirmed),
recovered_cum = cumsum(recovery),
death_cum = cumsum(death),
active_cum = cumsum(active)
)%>%
dplyr::mutate(recovered_cum= ifelse(date>as.Date("2021-08-04"), 191188,recovered_cum))
Tme <- nrow(df_daily)
t_start <- seq(2, Tme-35) # starting at 2 as conditional on the past observations
t_end <- t_start + 35 # adding 6 to get 7-day windows as bounds included in window
res_biweekly <- estimate_R(df_daily$confirmed,
method="parametric_si",
config = make_config(list(
t_start = t_start,
t_end = t_end,
mean_si = 4.8,
std_si = 2.3))
)
estimatedReff <- plot(res_biweekly, "R", color="red") + theme_bw() + labs(x="Time") +
ggplot2::scale_x_continuous(breaks=c(seq(1,tail(res_biweekly$dates,1), by=56)),labels=substr(seq.Date(as.Date("2020-03-10"), today()-1, by="8 weeks"),1,7))+theme(axis.text = element_text(face="bold", color="black", size=13))
ggplotly(estimatedReff)
r_eff<- data.frame(ro=c(2.4,res_biweekly$R$`Quantile.0.975(R)`), date=c(0,res_biweekly$R$t_end))
mu <- 4.8 # mean in days days
sigma <- 2.3 # standard deviation in days # https://www.ijidonline.com/article/S1201-9712(20)30119-3/pdf
#ggsave("estimatedR.png", estimatedReff, width=16, height=10)
```
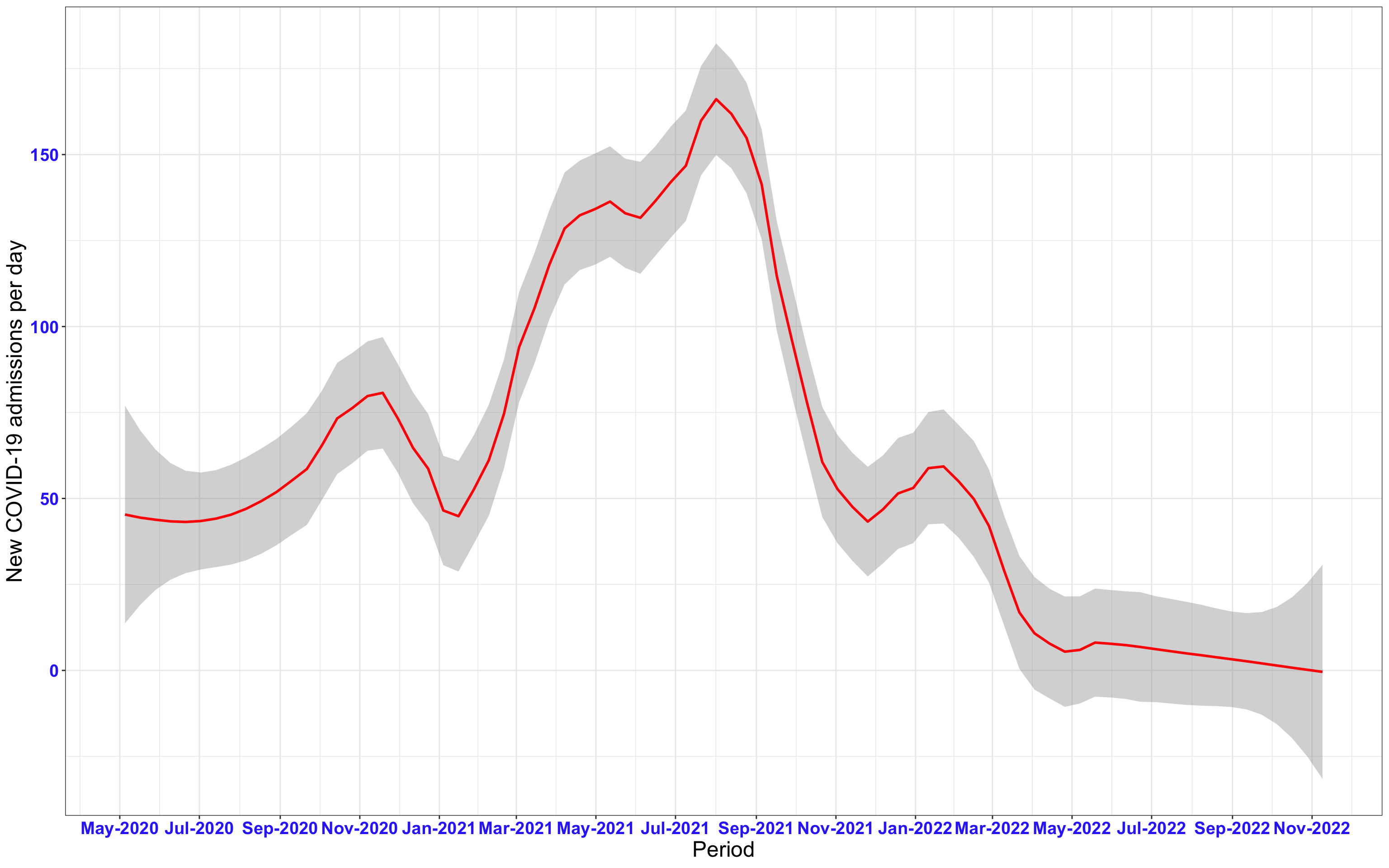
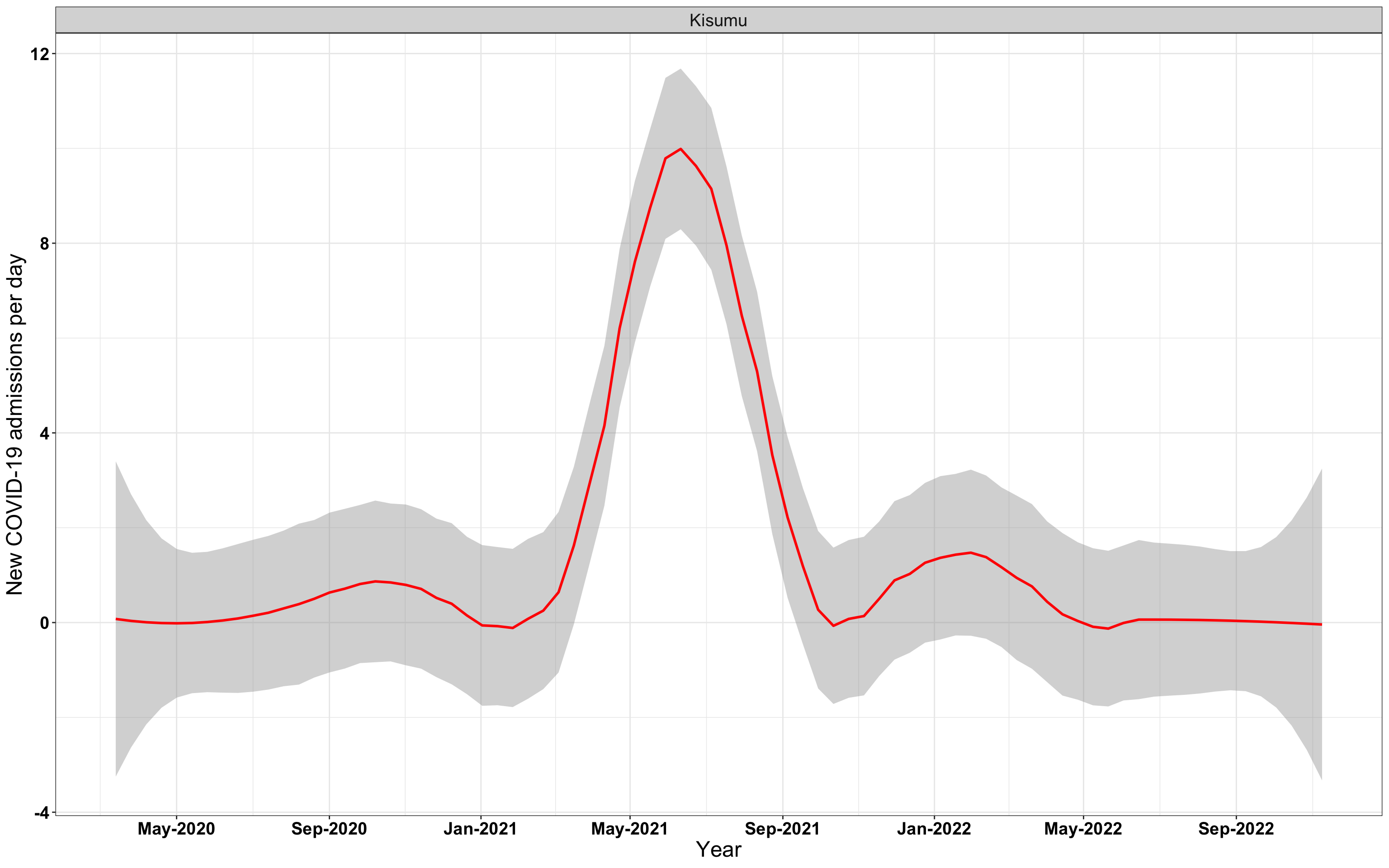
# Kenya {data-navmenu="Hospital admissions"}
```{r,fig.width=16, fig.height=10}
data<-read_csv("admissions_summary_20221109.csv")%>%
mutate_at(vars(admissions, new_admissions, icu, oxygen, new_discharges), funs(abs(.)))%>%
mutate(date=as.Date(date, "%d/%m/%Y"))%>%
filter(date<(today()+1))
data1<- data%>%
filter(date!=as.Date("2021-03-23"))%>%
group_by(date)%>%
mutate(admissions=sum(admissions))%>%
mutate(new_admissions=sum(new_admissions))%>%
mutate(icu=sum(icu))%>%
mutate(oxygen=sum(oxygen))%>%
mutate(new_discharges=sum(new_discharges))%>%
select(-county)%>%
ungroup()%>%
distinct()
data_county<- data%>%
dplyr::group_by(date, county)%>%
mutate(admissions=sum(admissions))%>%
mutate(new_admissions=sum(new_admissions))%>%
mutate(icu=sum(icu))%>%
mutate(oxygen=sum(oxygen))%>%
mutate(new_discharges=sum(new_discharges))%>%
ungroup()%>%
distinct()#%>%
#filter(county%in%c("Uasin Gishu", "Murang'a", "Laikipia", "Siaya", "Nyeri", "Nyamira", "Kitui") & date>as.Date("2021-03-14"))
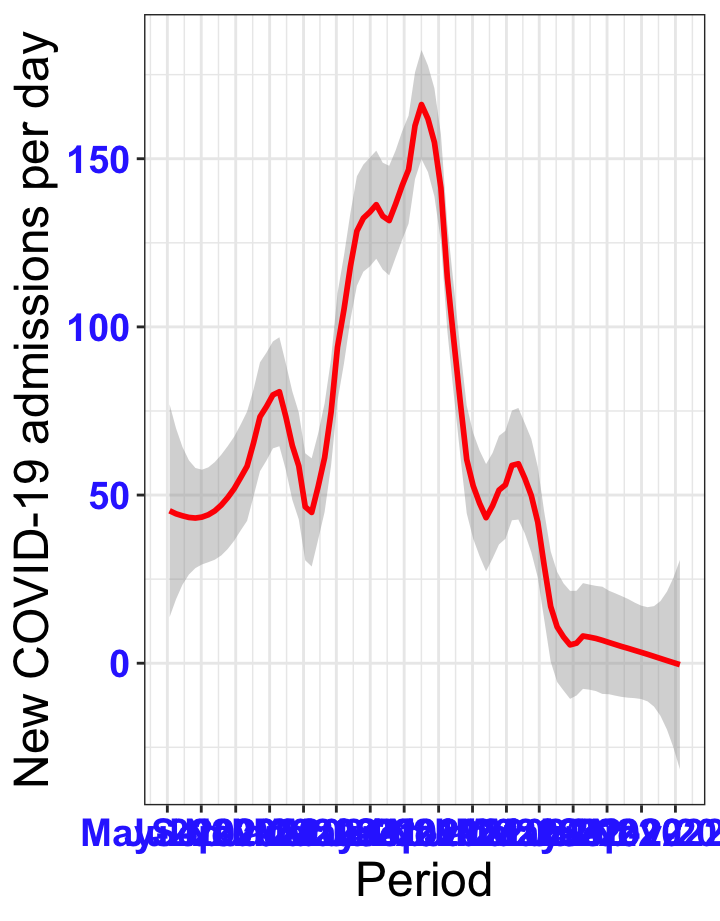
p1<-ggplot(data1[data1$date>as.Date("2020-05-04"),], aes(x=date,y=new_admissions))+theme_bw()+geom_smooth(span=0.3, colour="red", oob="squish")+scale_x_date(date_labels ="%b-%Y" ,date_breaks = "2 months")+labs(x="Period", y="New COVID-19 admissions per day")+theme(text=element_text(size=18 ), axis.text = element_text(color = "#3339FF", face = "bold"))
x<-unique(data_county$county)
p1
# ggplotly(p1)%>%
# partial_bundle()
```
```{r,include=F}
xmax<-max(data_county$date)-8
data_county1<- data_county%>%
filter(date>xmax)%>%
group_by(county)%>%
mutate(icu=round(mean(icu)))%>%
mutate(new_admissions=round(mean(new_admissions)))%>%
ungroup()%>%
select(-oxygen, -new_discharges, -admissions, -date)%>%
distinct()%>%
mutate(county=recode(county,"Murang'a"="Muranga"))
county<-st_read("County.shp")
county_admissions<-full_join(county, data_county1, by=c("Name"="county"))
plot1<-ggplot(county_admissions, aes(fill=new_admissions, label=Name))+geom_sf()+theme_void()+scale_fill_gradient(high="#f03b20", low="#ffeda0", na.value = "grey80")+labs(fill="",title="Average number of hospital admissions over the last one week")
```
# Hospital admissions {data-navmenu="Visualisation"}
```{r,fig.width=16, fig.height=10, message=F}
plotly::ggplotly(plot1)
```
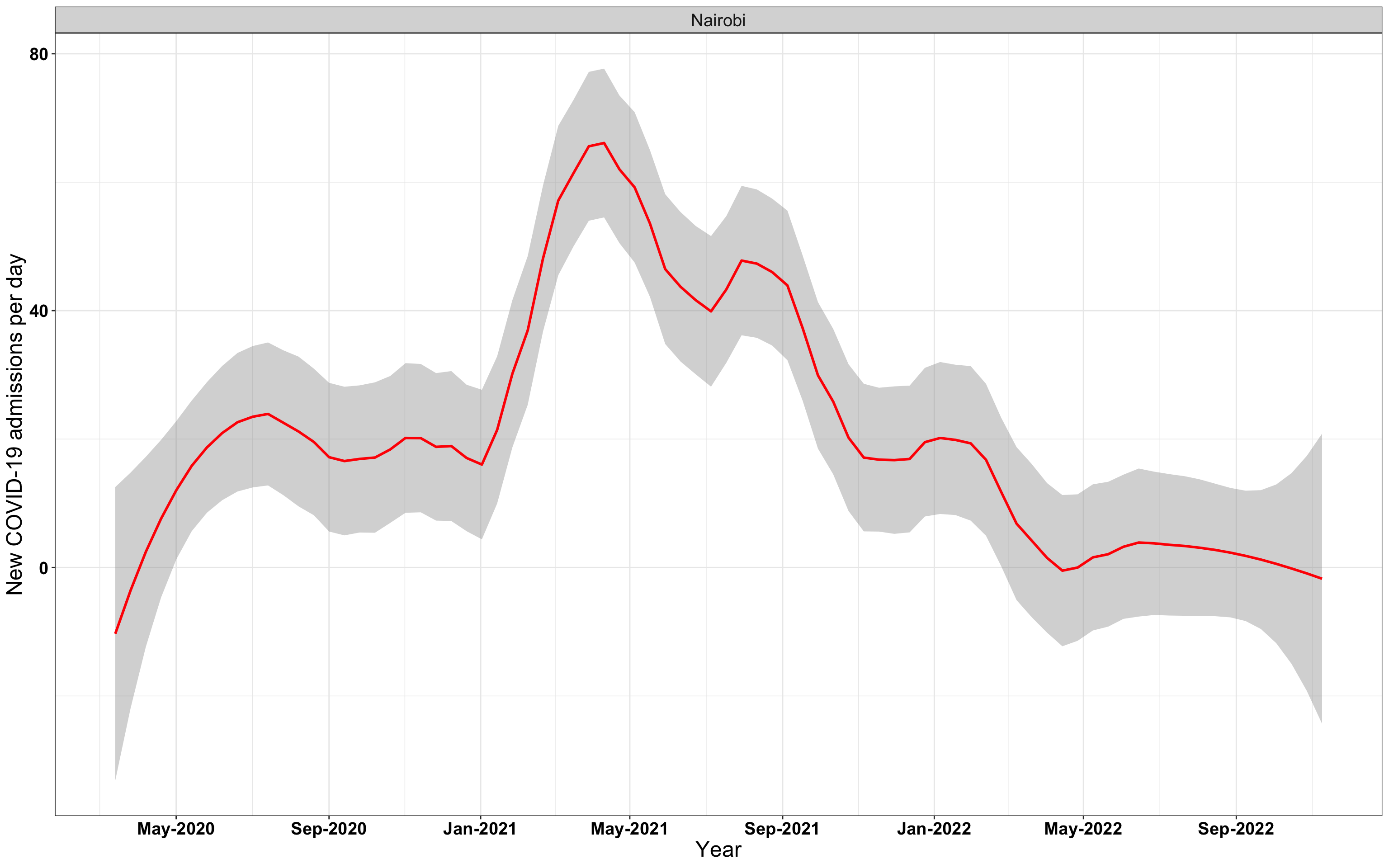
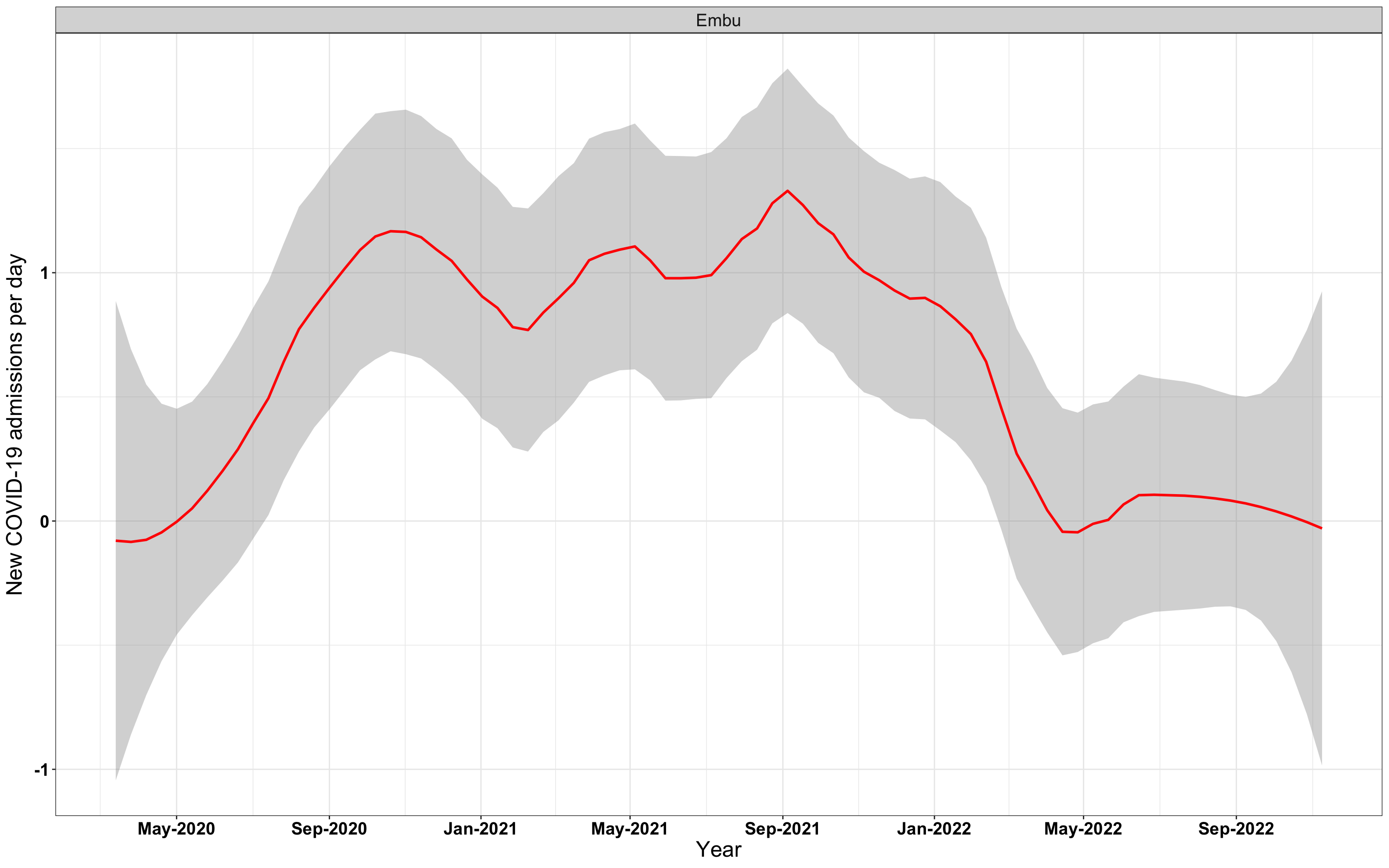
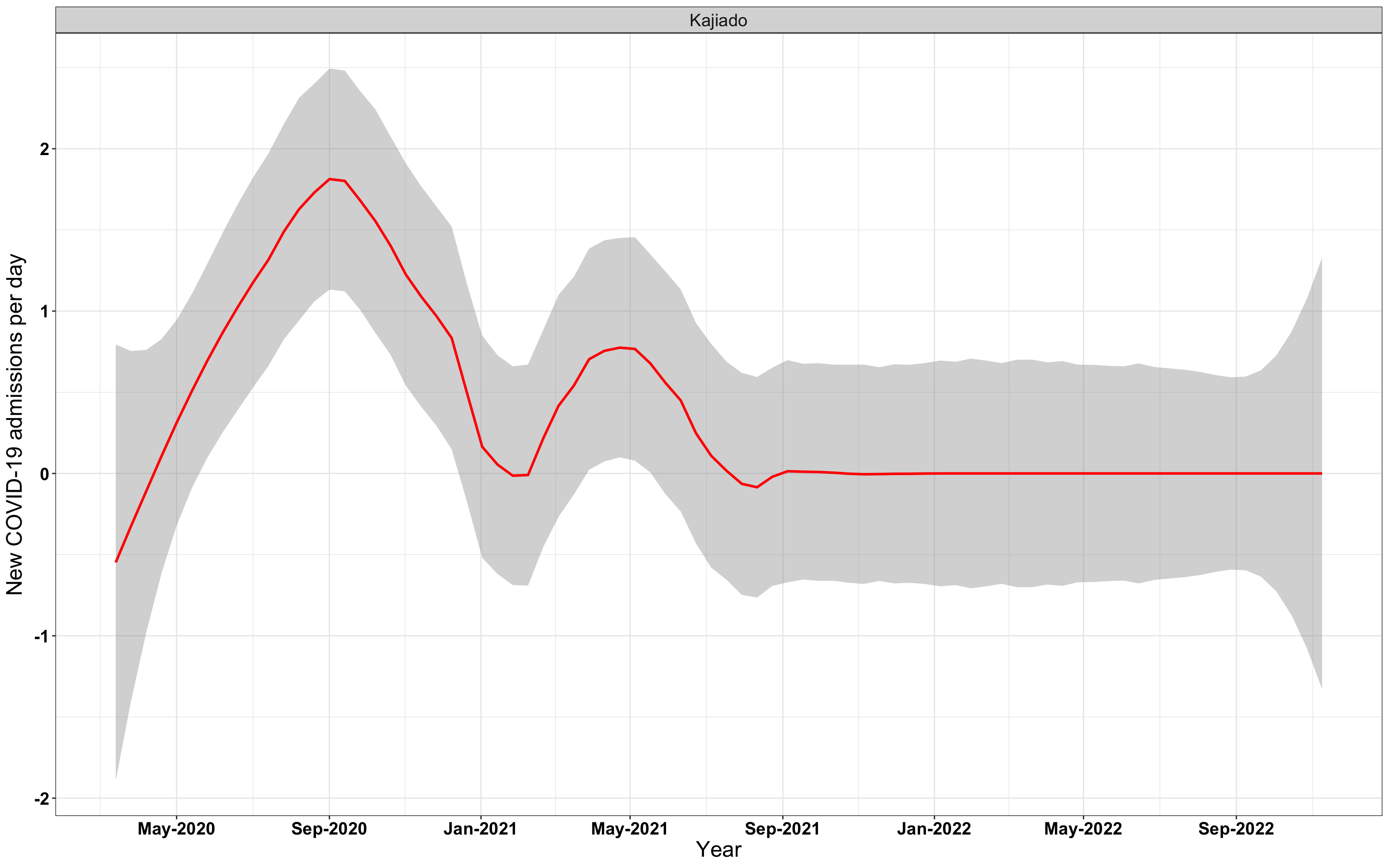
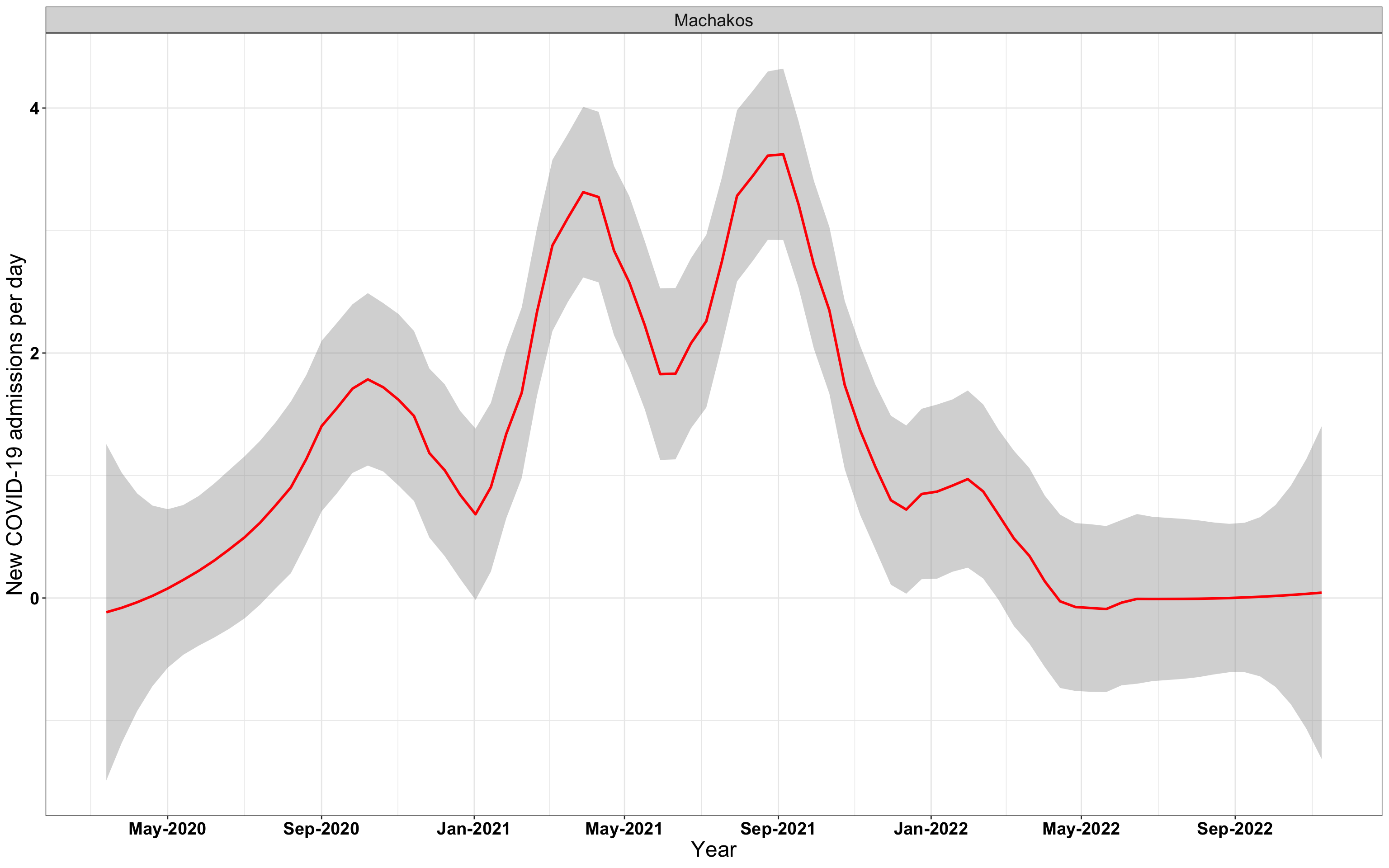
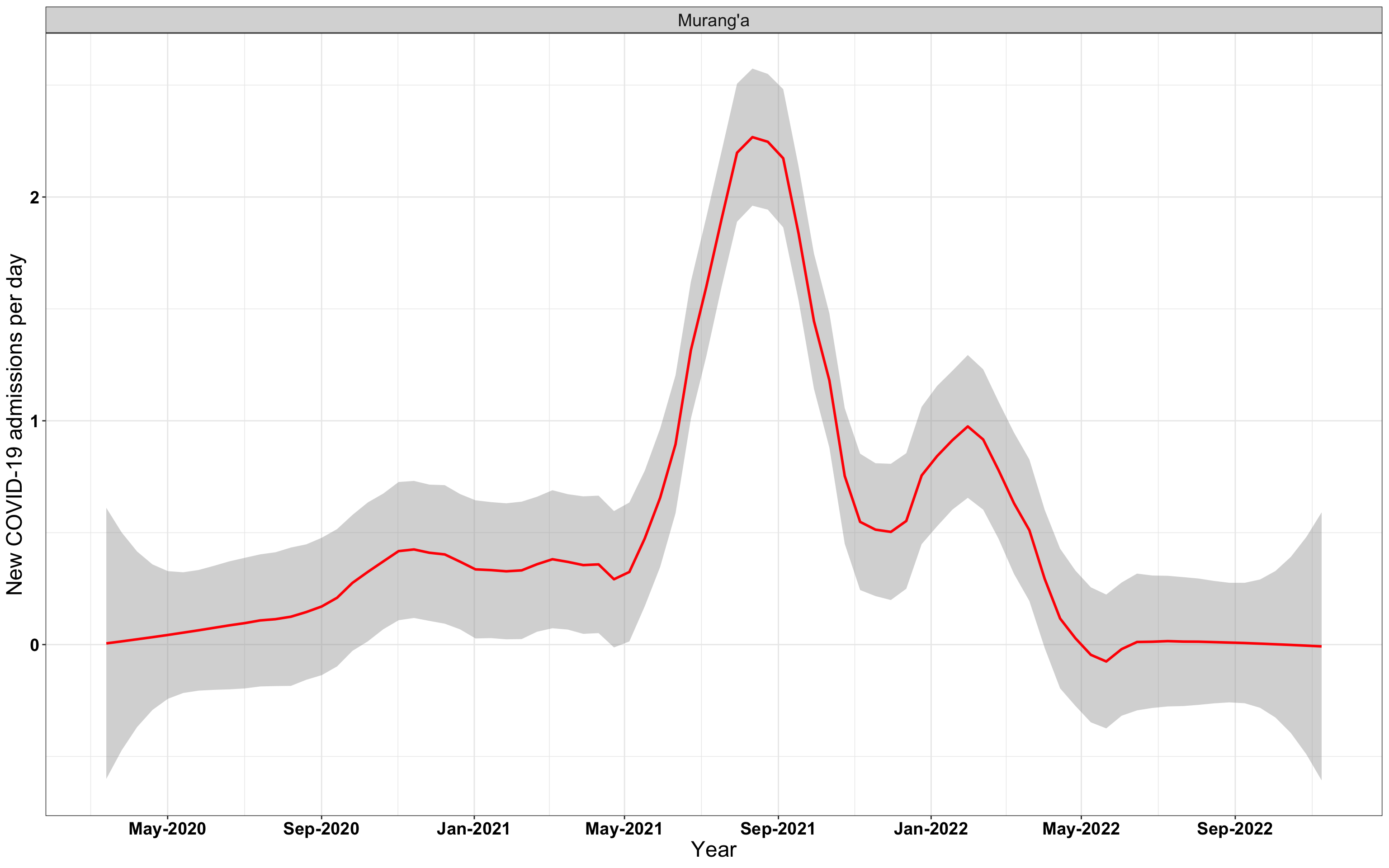
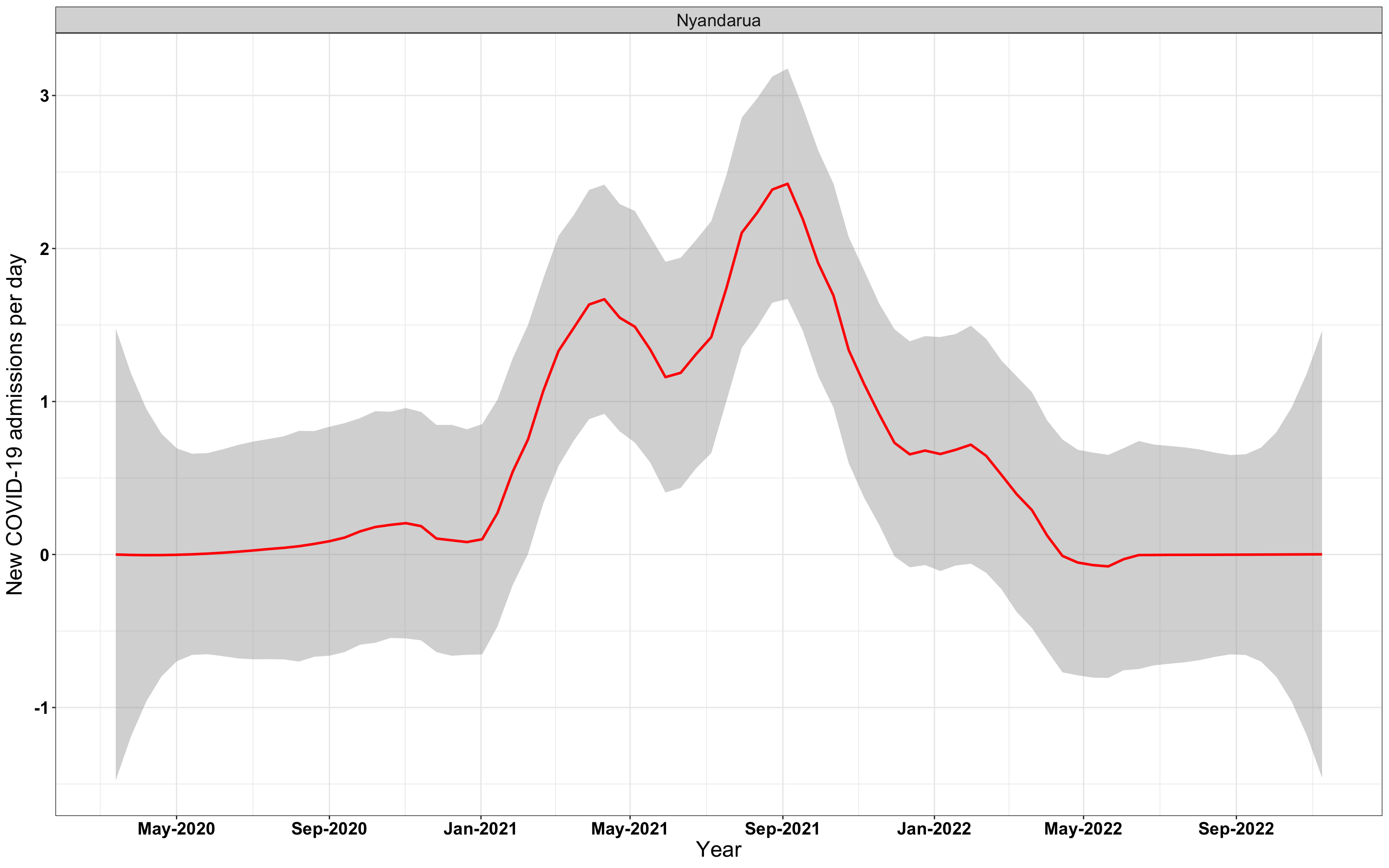
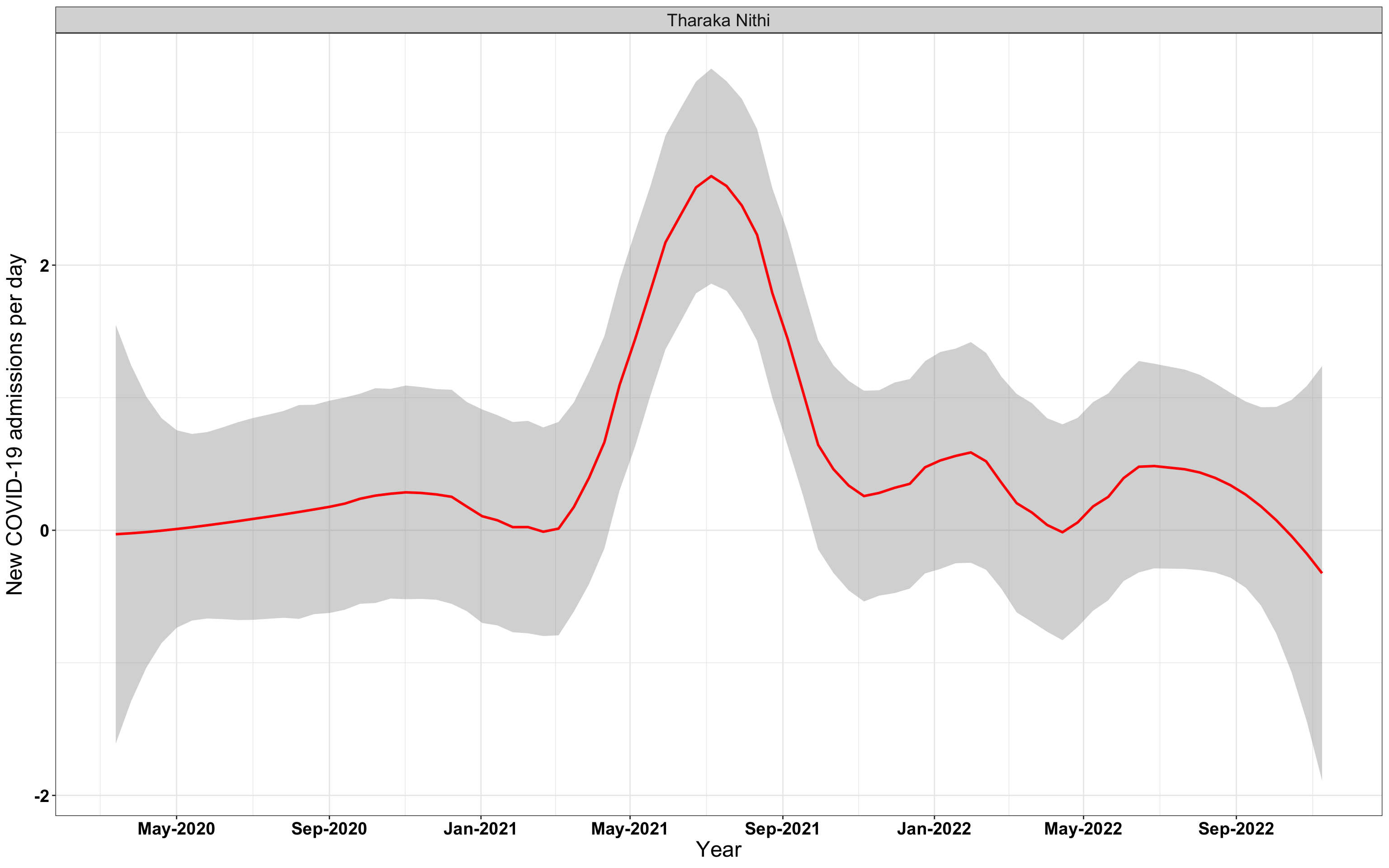
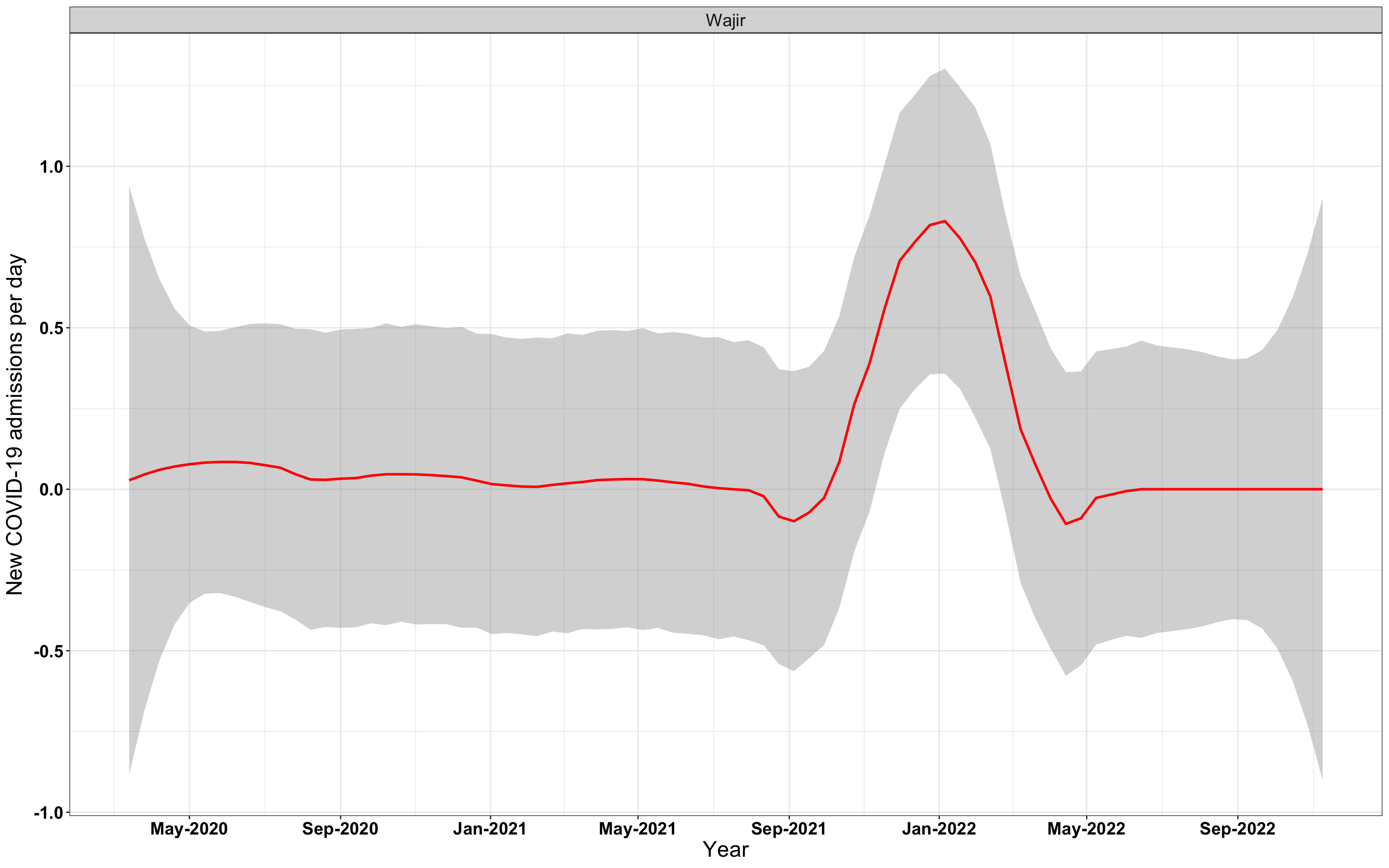
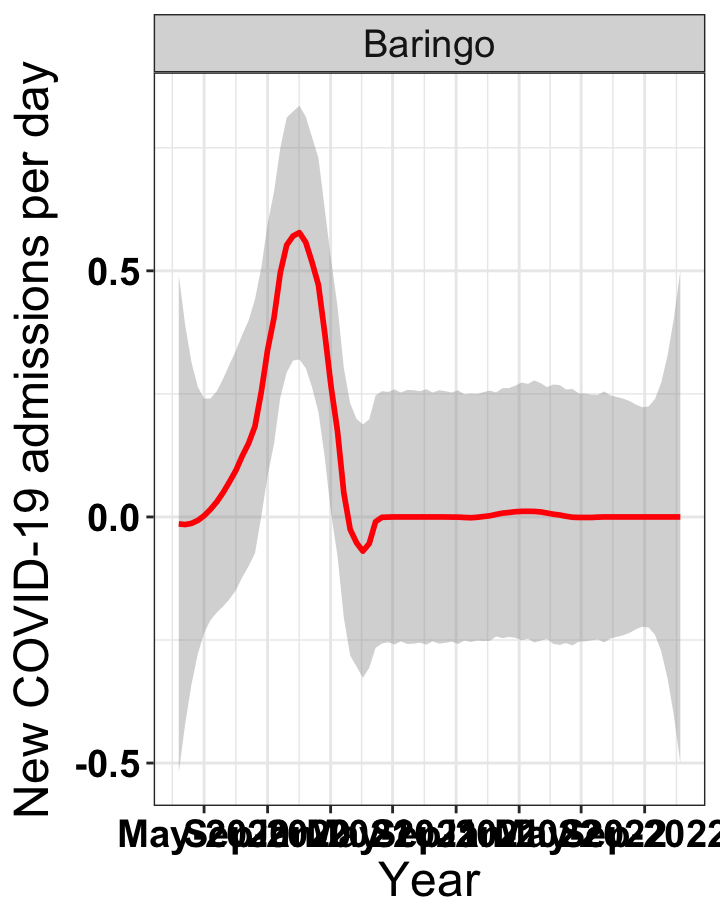
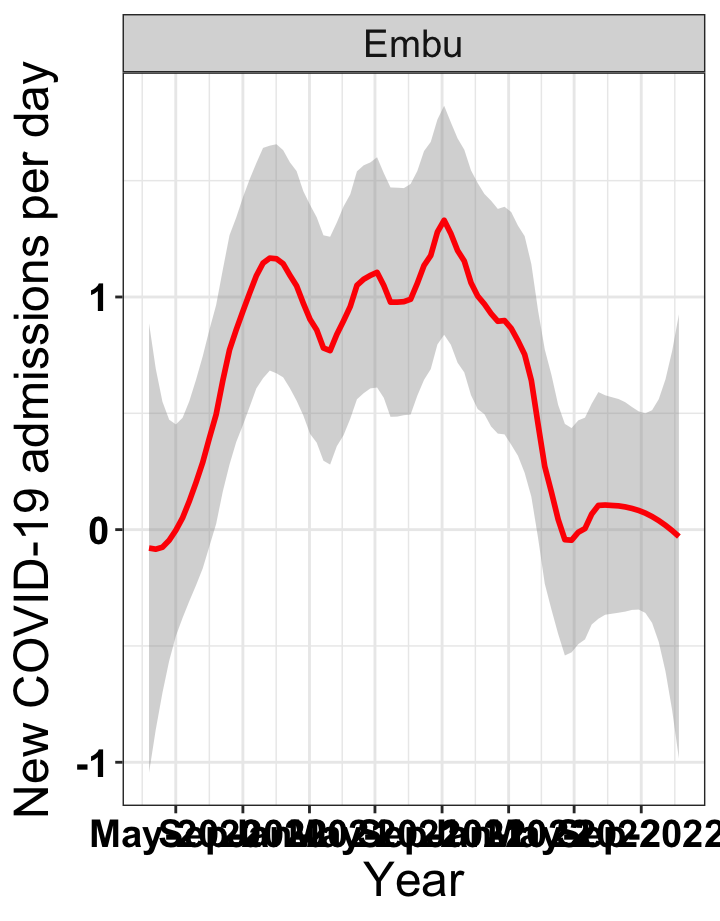
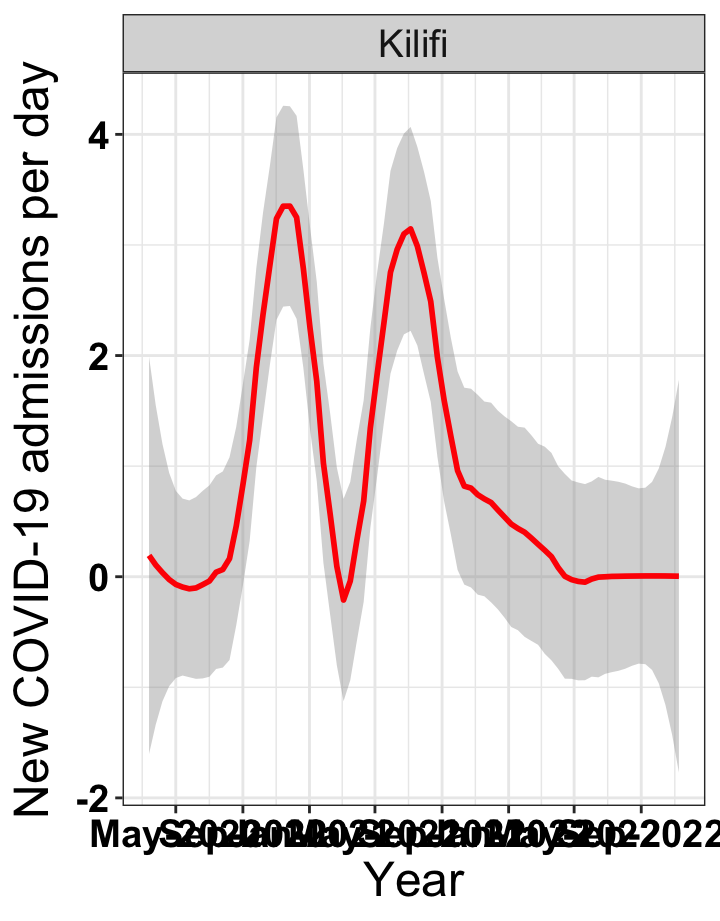
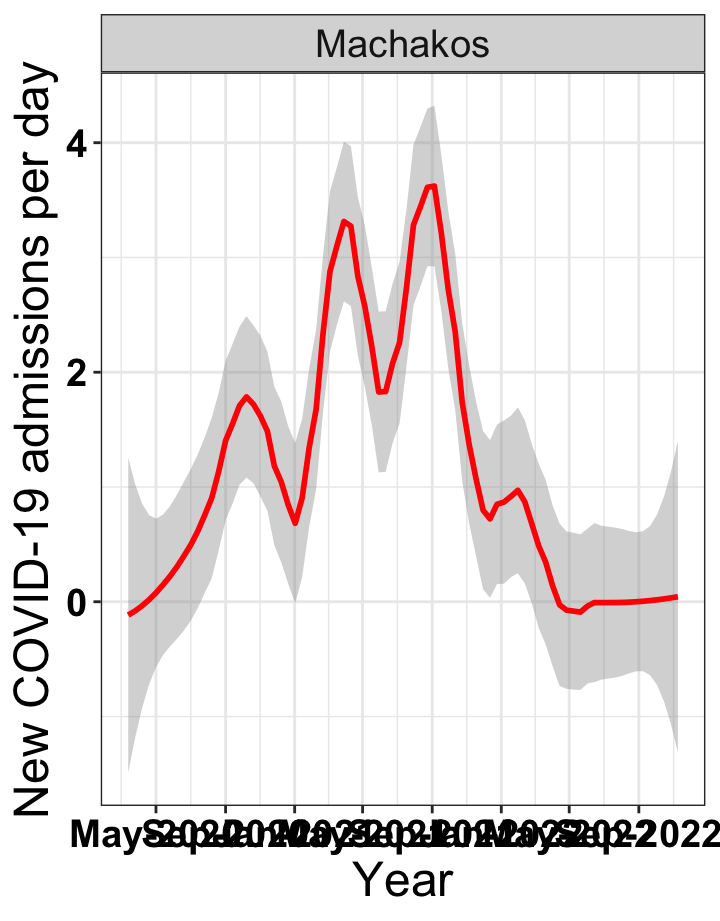
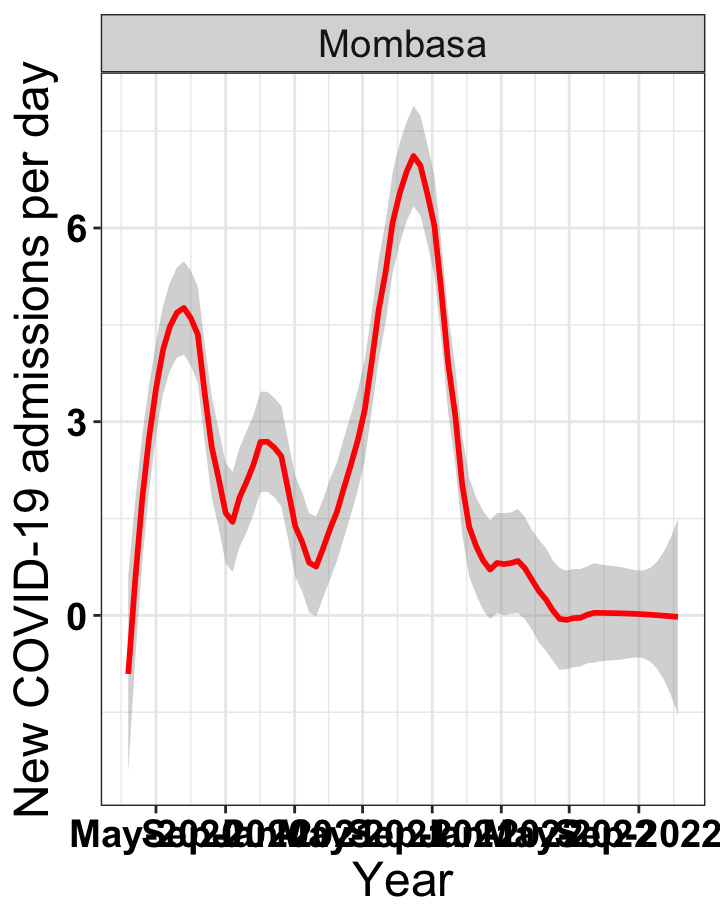
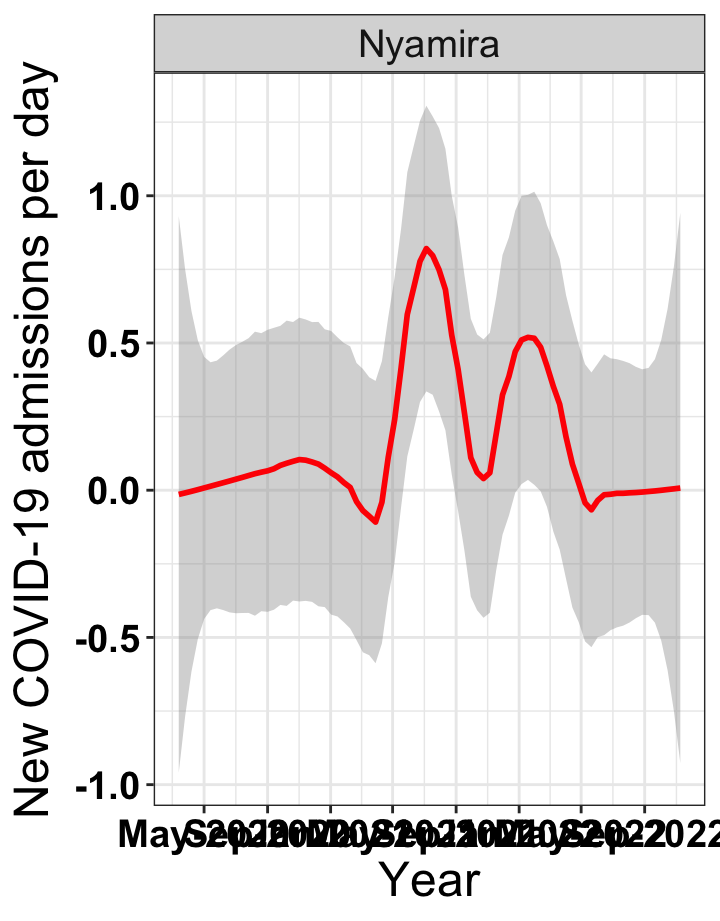
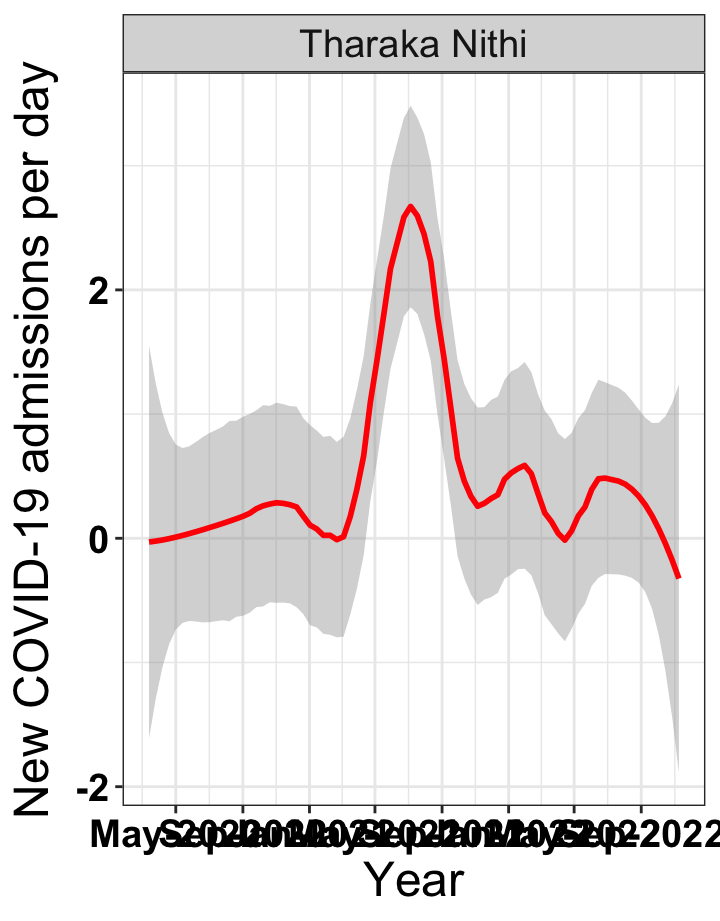
# `r x[1]` {data-navmenu="Hospital admissions"}
```{r,fig.width=16, fig.height=10, results="asis"}
data_county<-data_county%>%
dplyr::filter(!is.na(county))%>%
mutate(date1=as.Date(date))%>%
filter(date>"2020-03-12")
library(pander)
graphs<-list()
for (i in 1:47) {
# Inserts Month titles
#pandoc.header(paste("{data-navmenu="Hospital admissions"}",x[1]), level = 1, "setex")
# Section contents
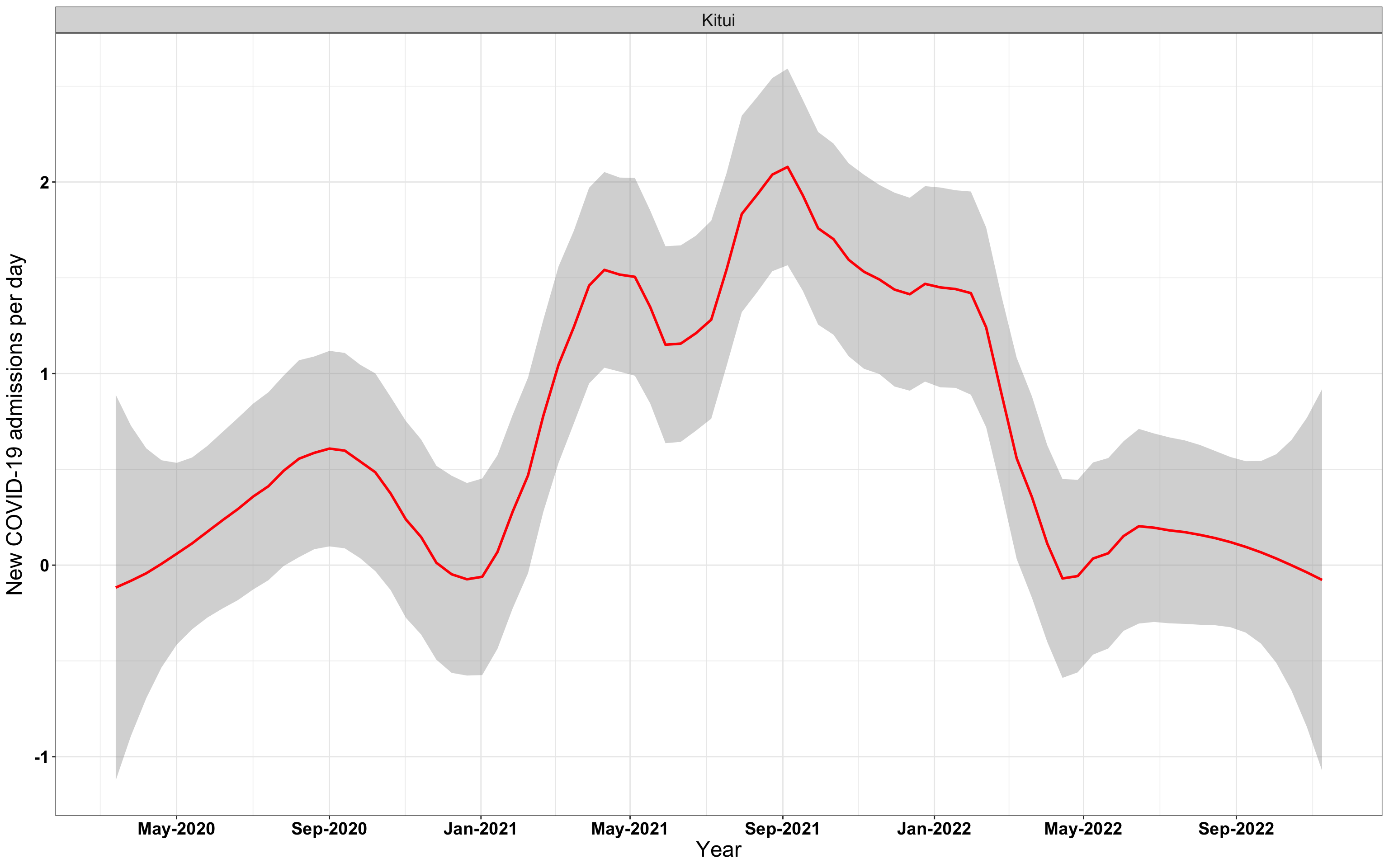
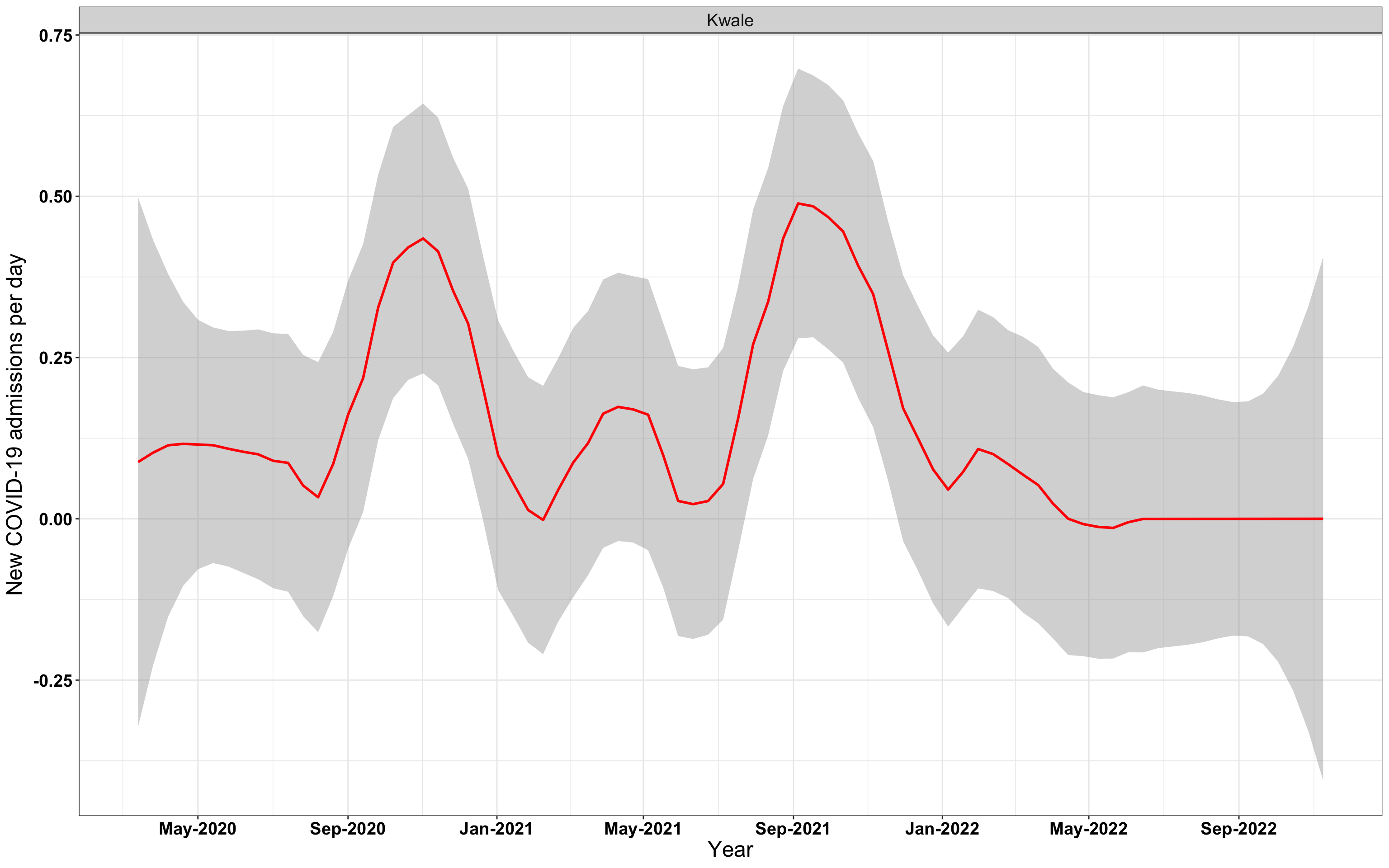
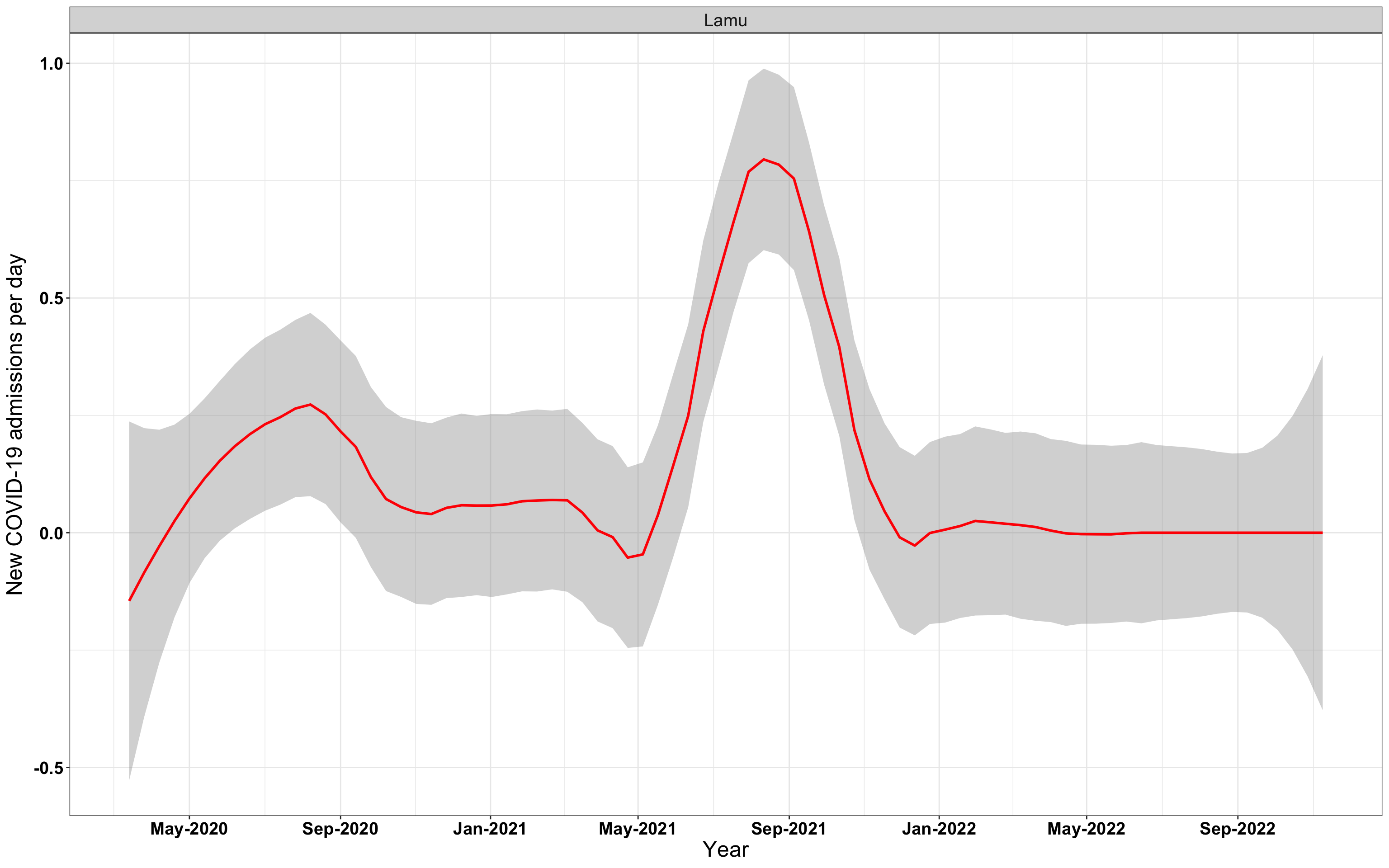
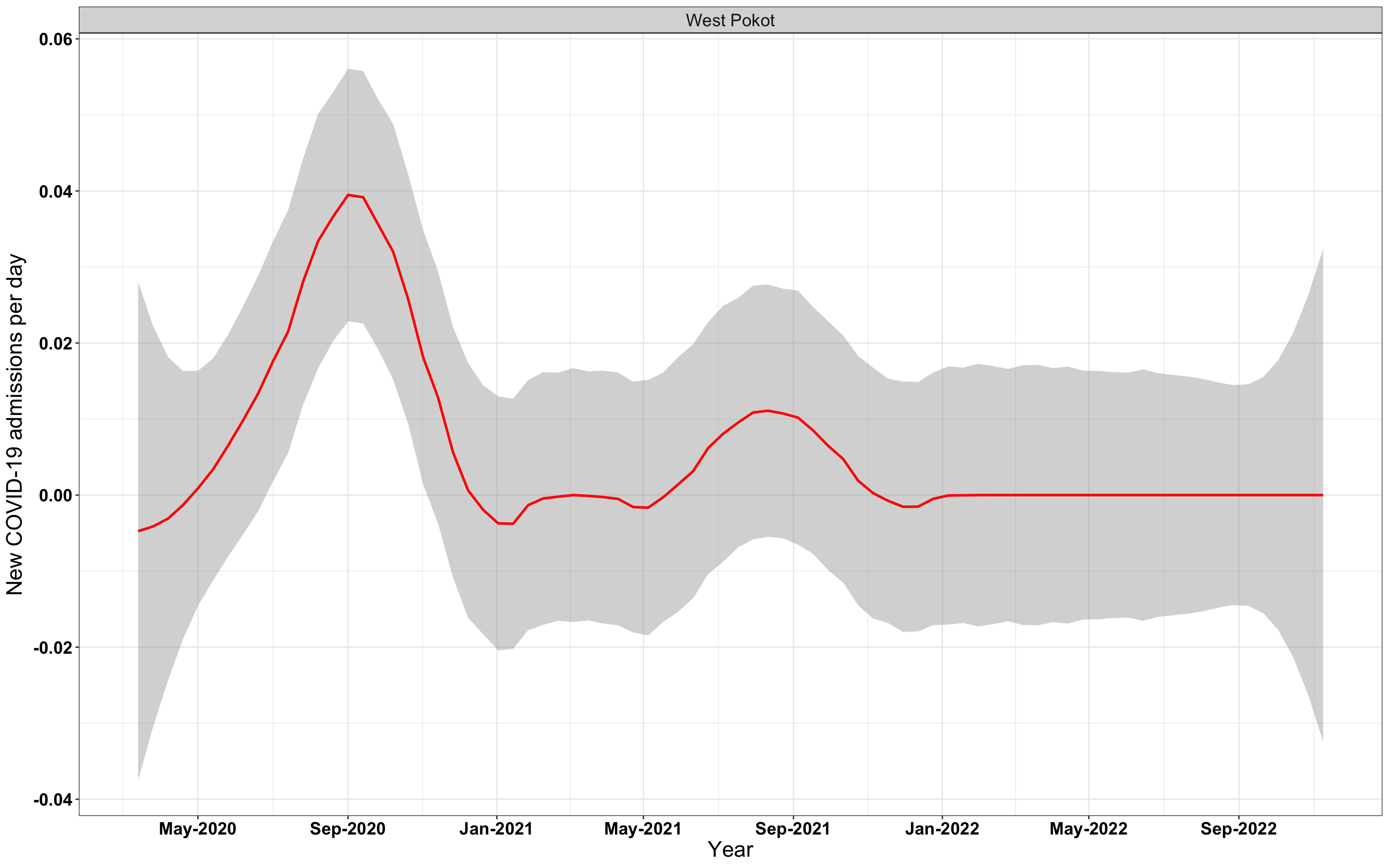
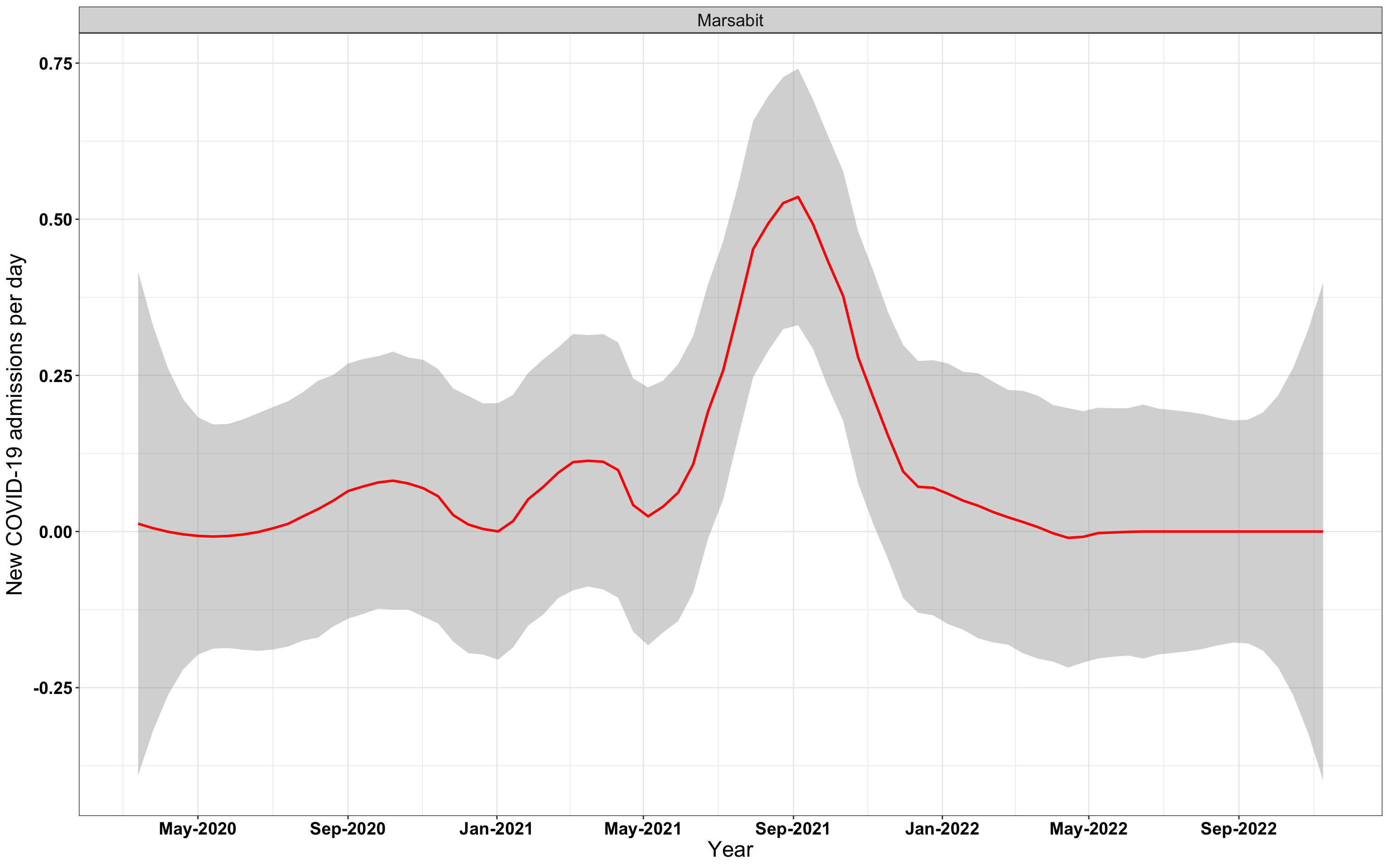
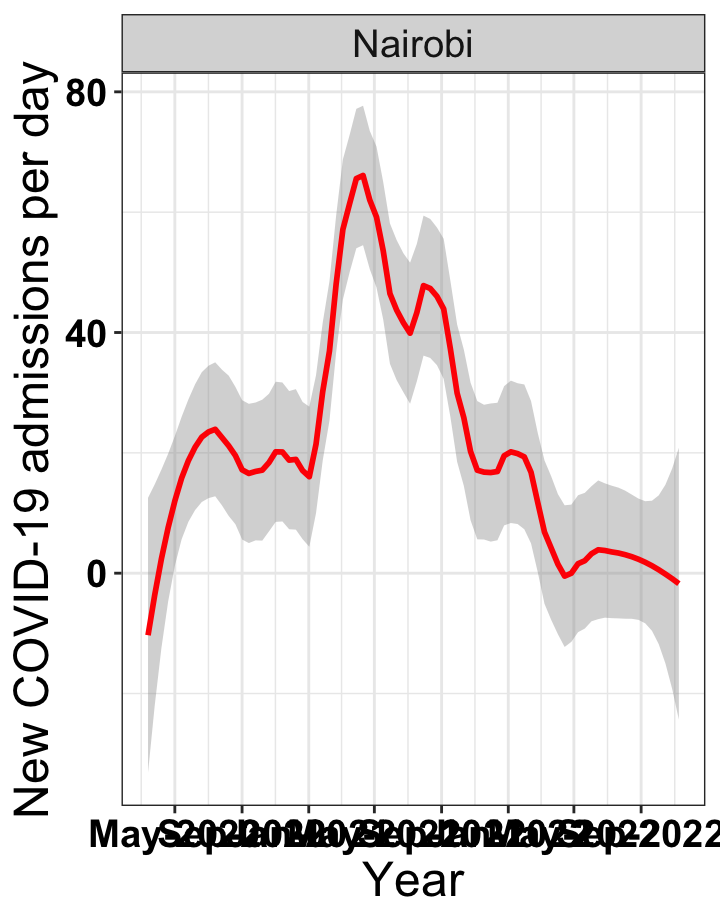
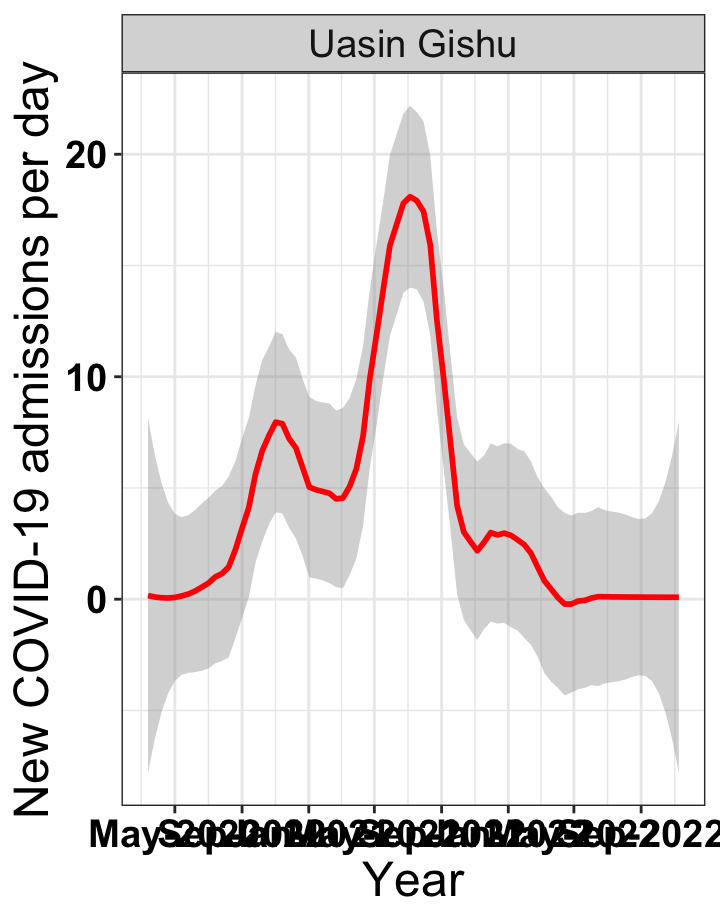
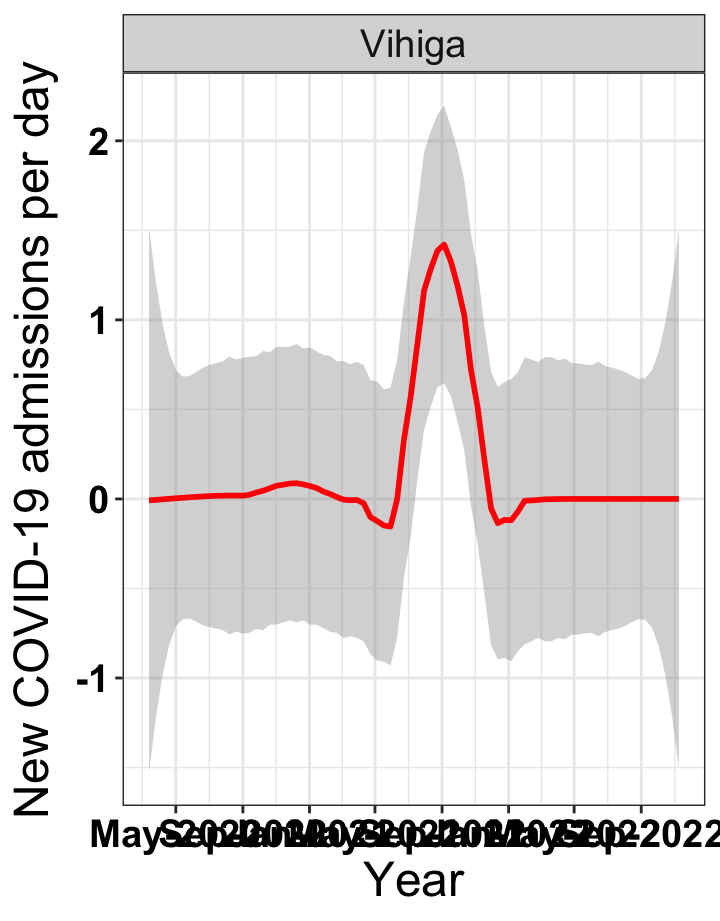
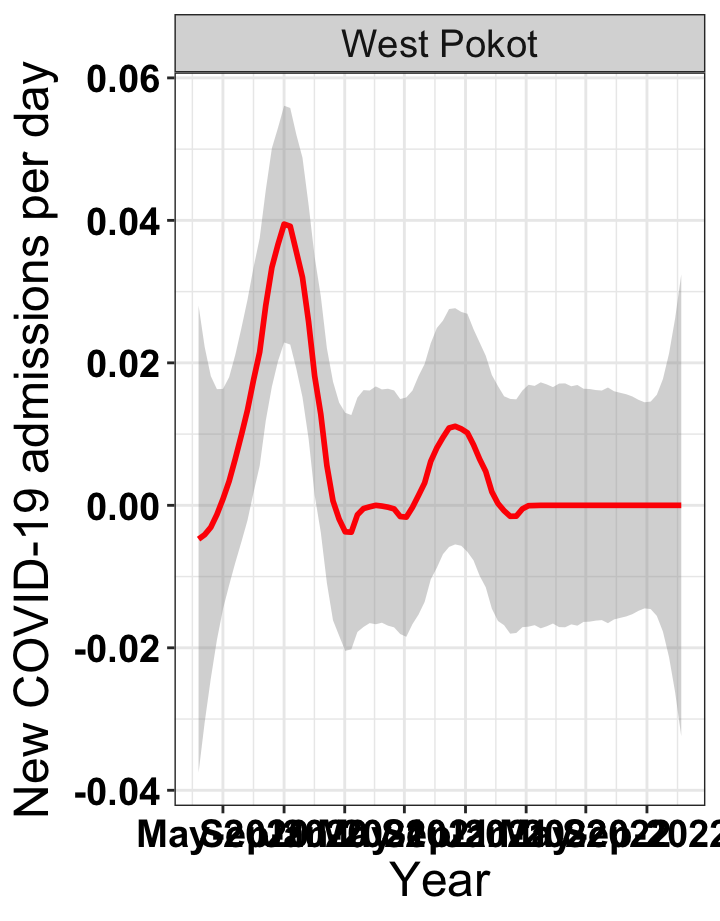
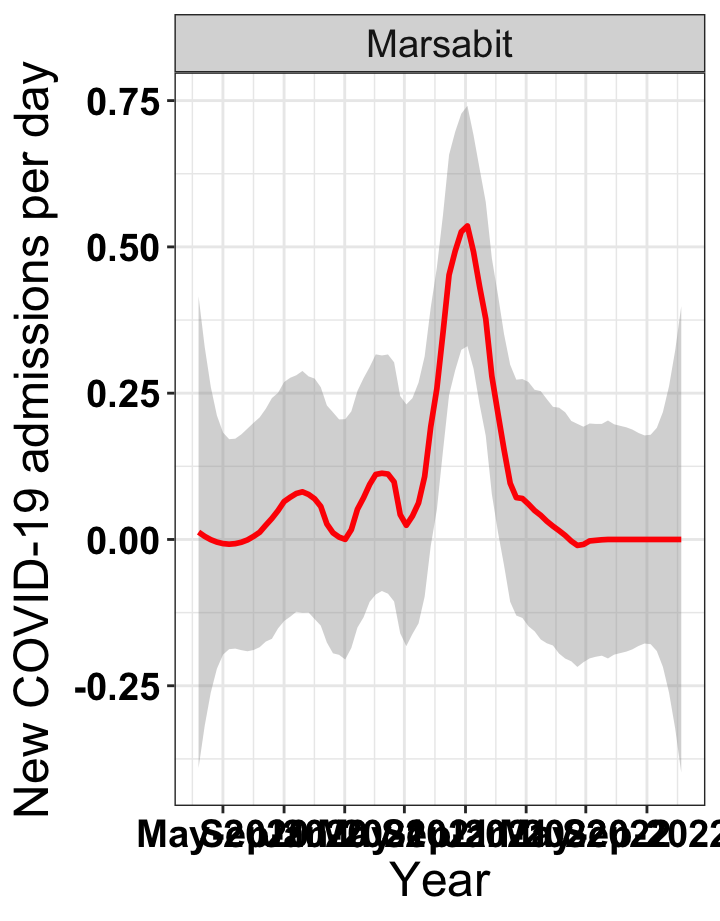
p1<-eval(substitute(ggplot(data_county[data_county$county%in%x[i],],aes(x=date1,y=new_admissions))+theme_bw()+facet_wrap(~county)+geom_smooth(span=0.3, colour="red", oob="squish")+labs(x="Year", y="New COVID-19 admissions per day")+theme(text=element_text(size=18 ), axis.text = element_text(color = "black", face = "bold"))+scale_x_date(breaks="4 months", date_labels = "%b-%Y"),list(i = i)))
graphs[[i]]<-p1
# adding also empty lines, to be sure that this is valid Markdown
# pander::pandoc.p('')
# pander::pandoc.p('')
}
graphs[[1]]
```
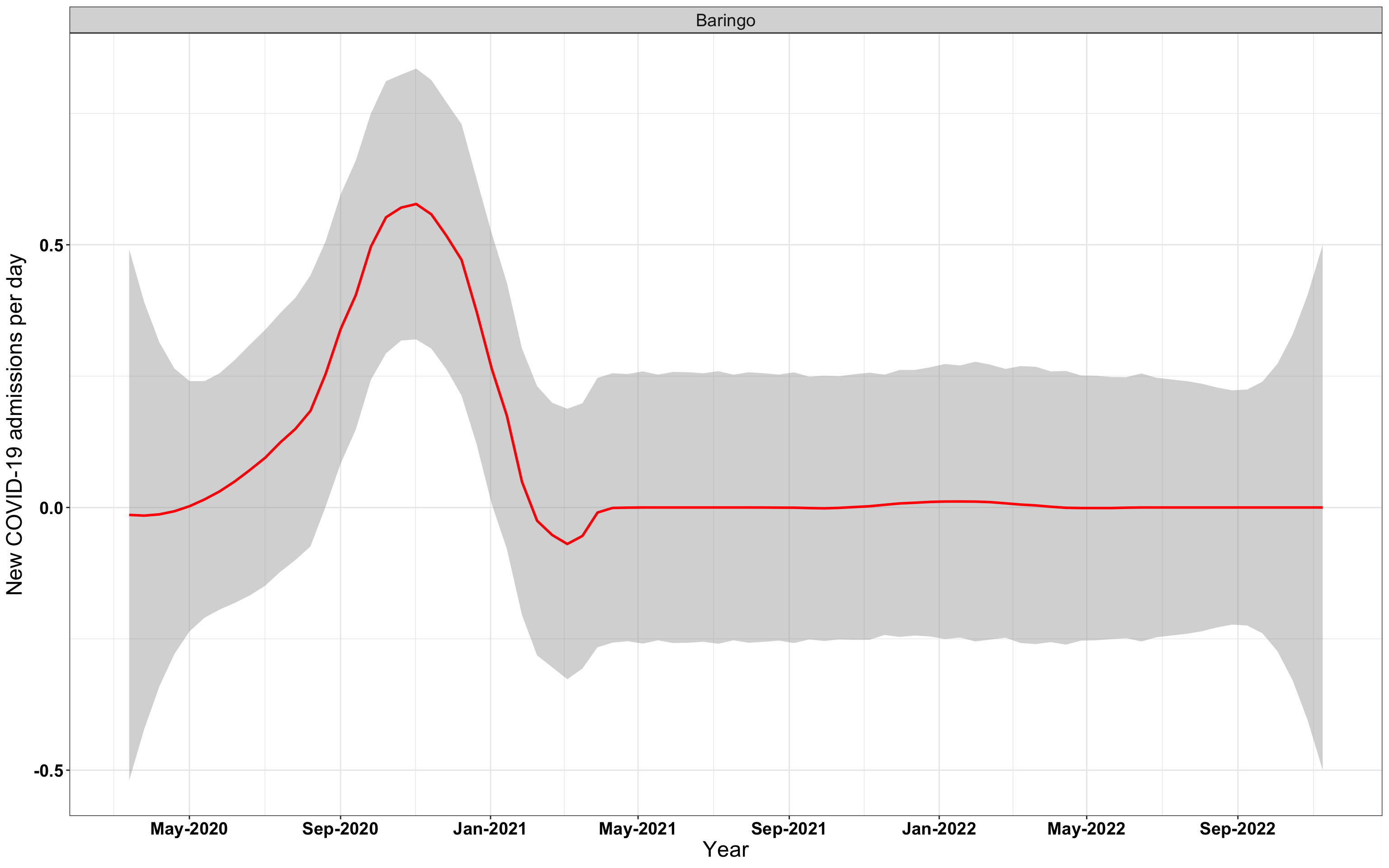
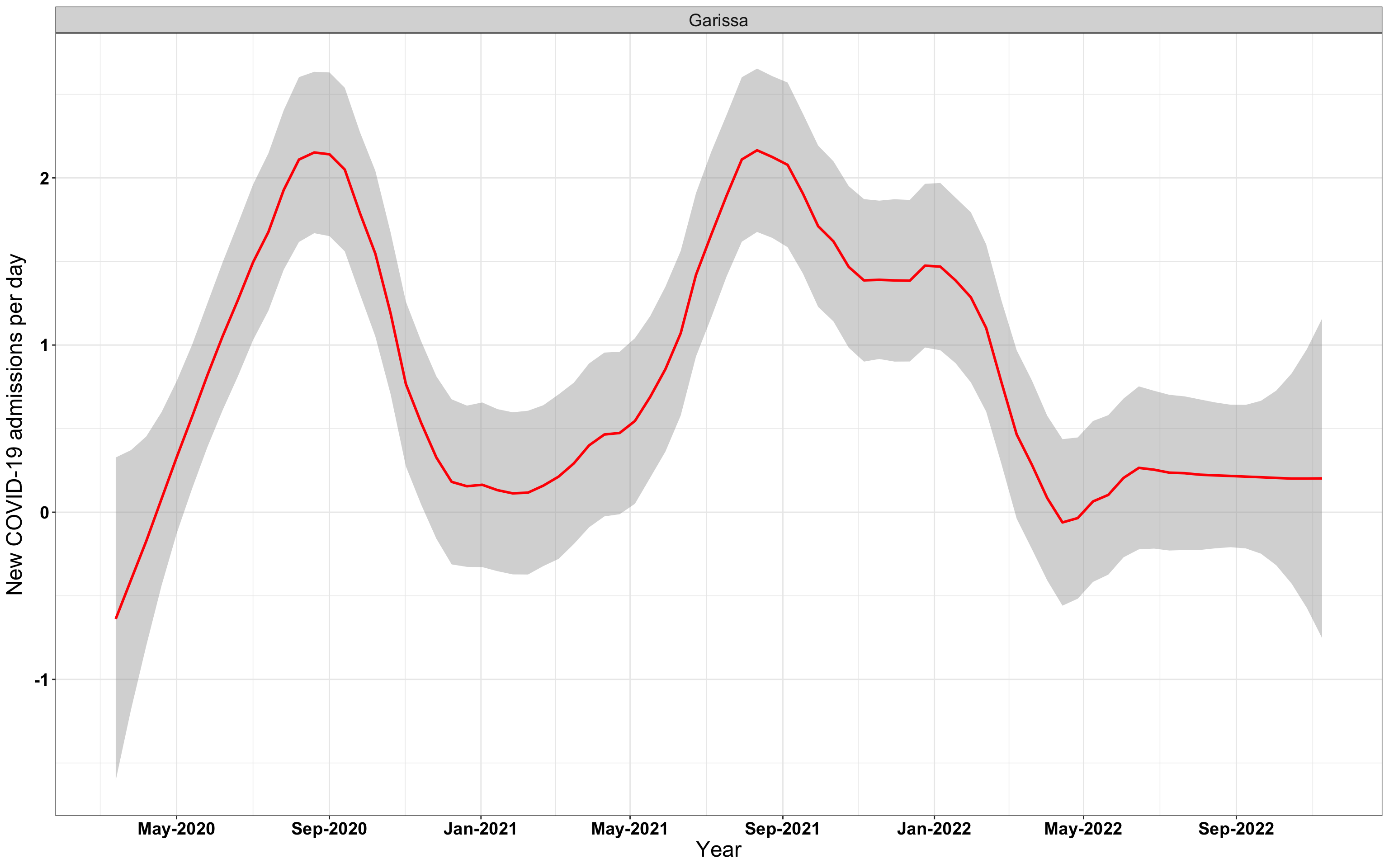
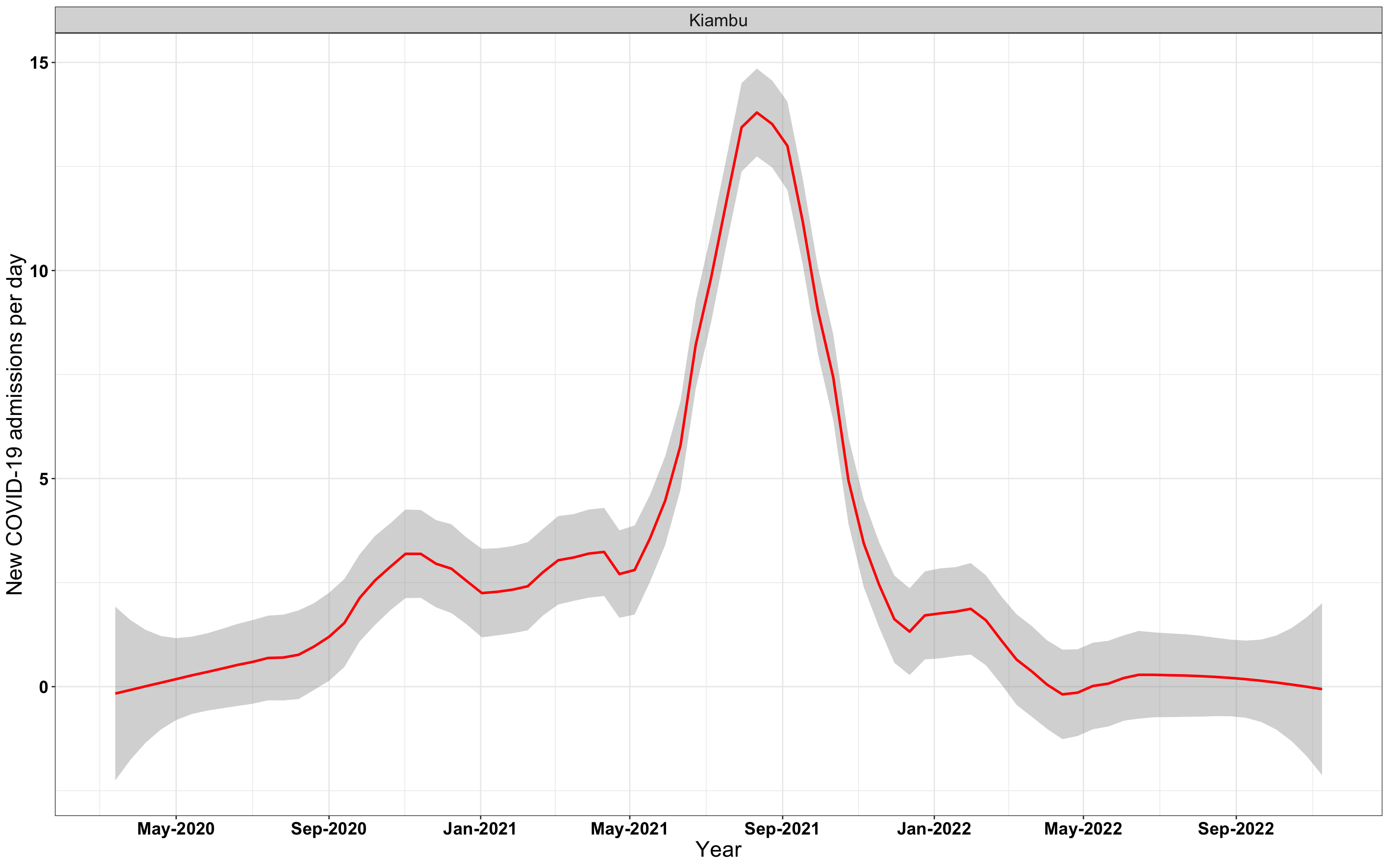
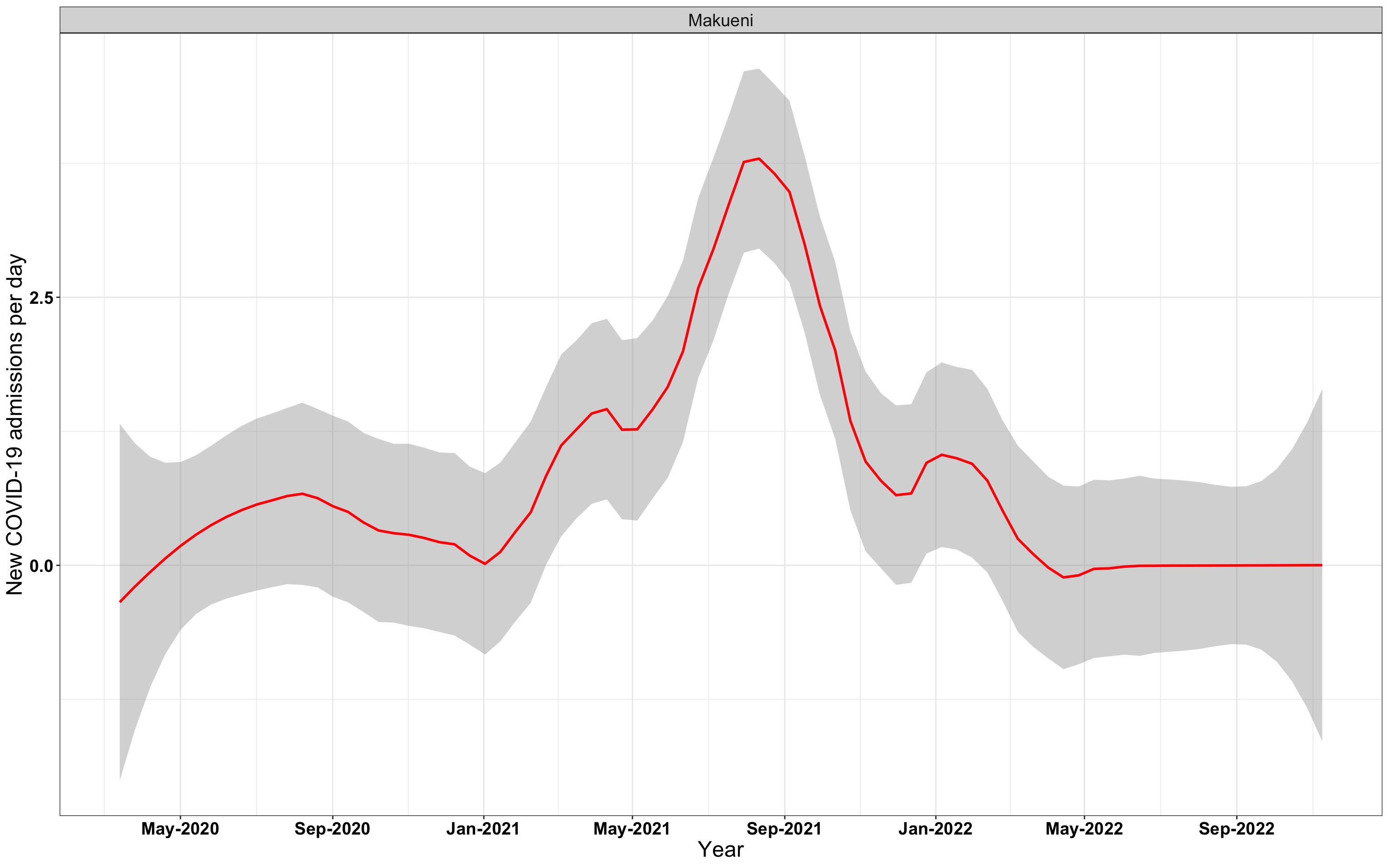
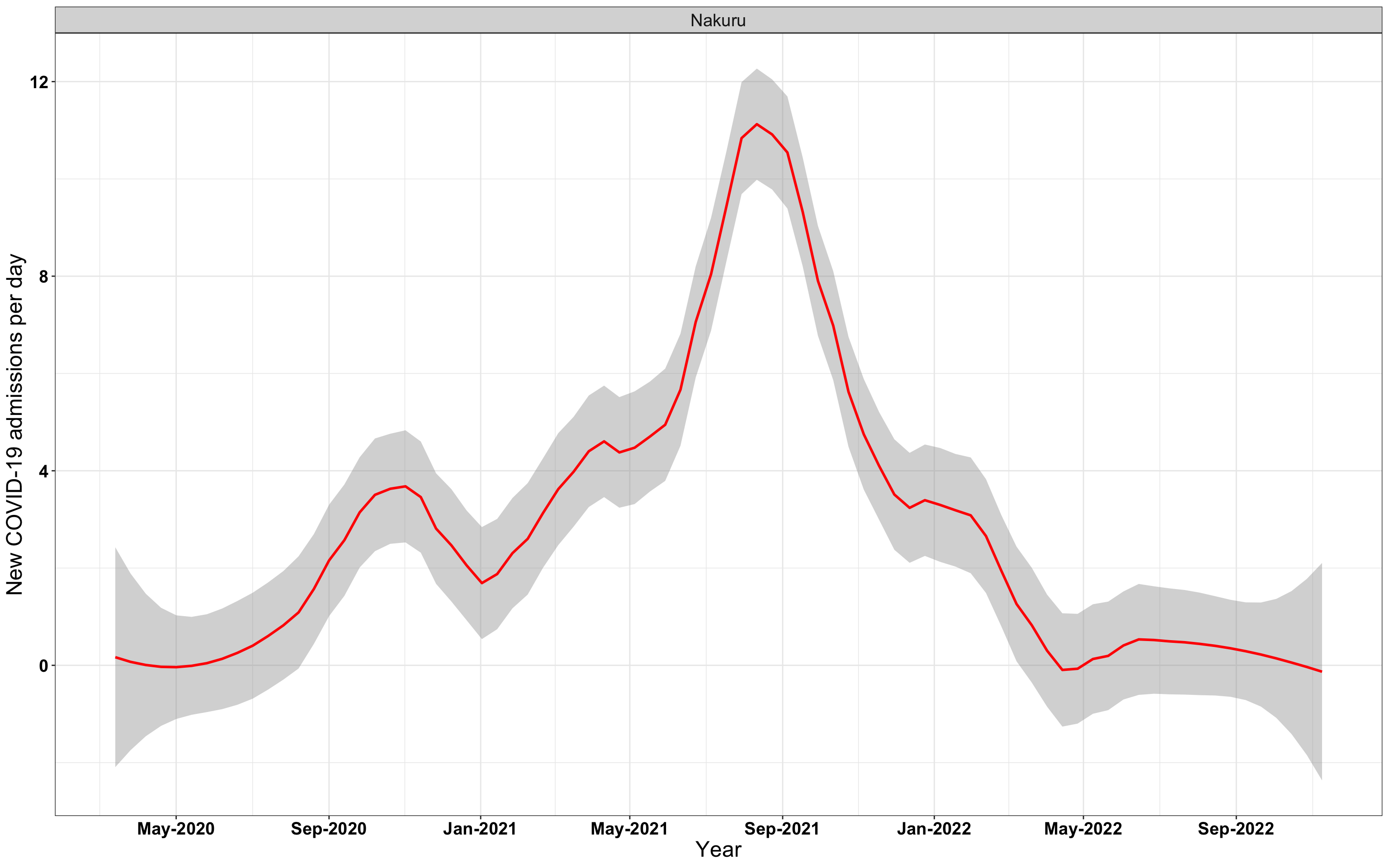
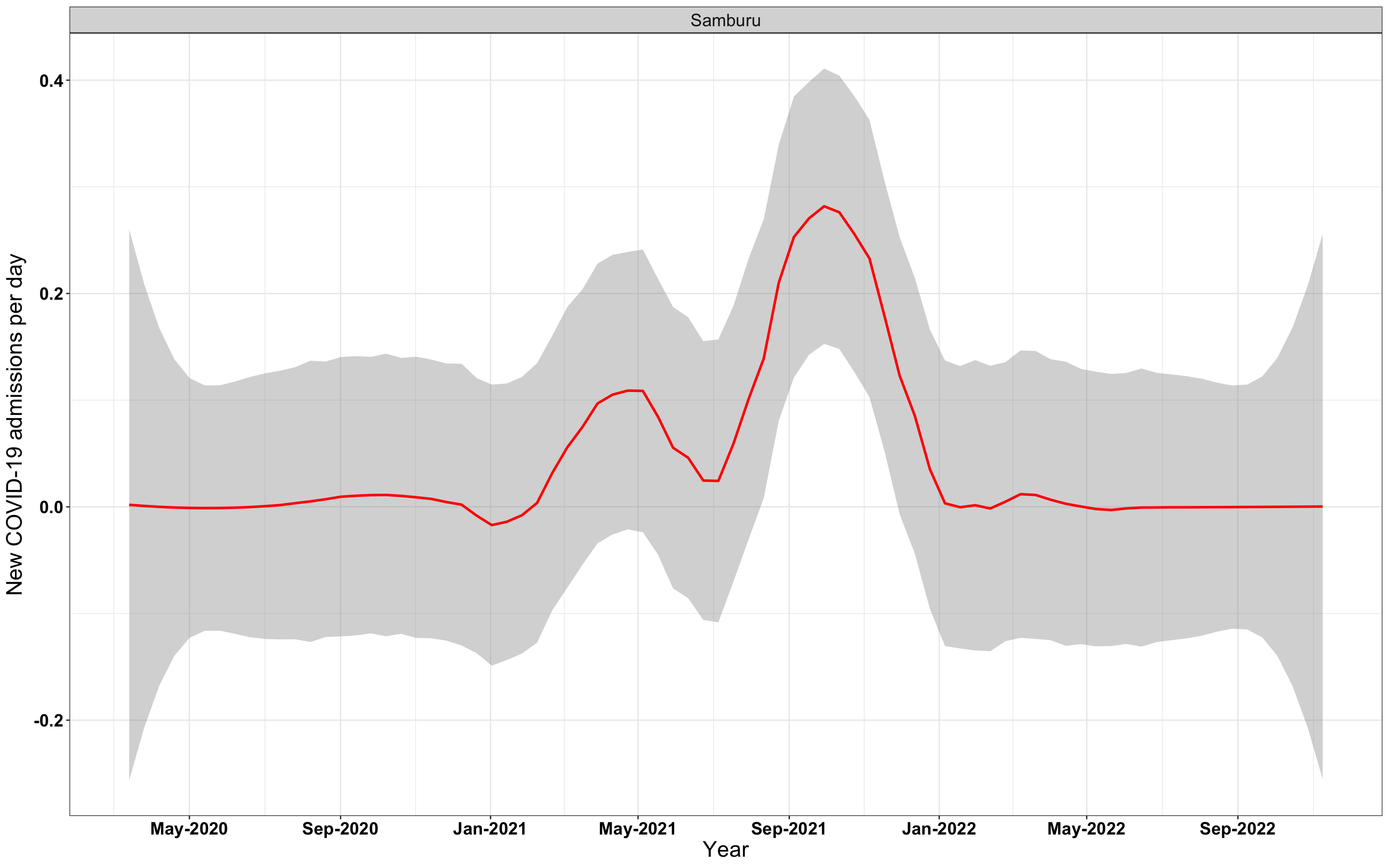
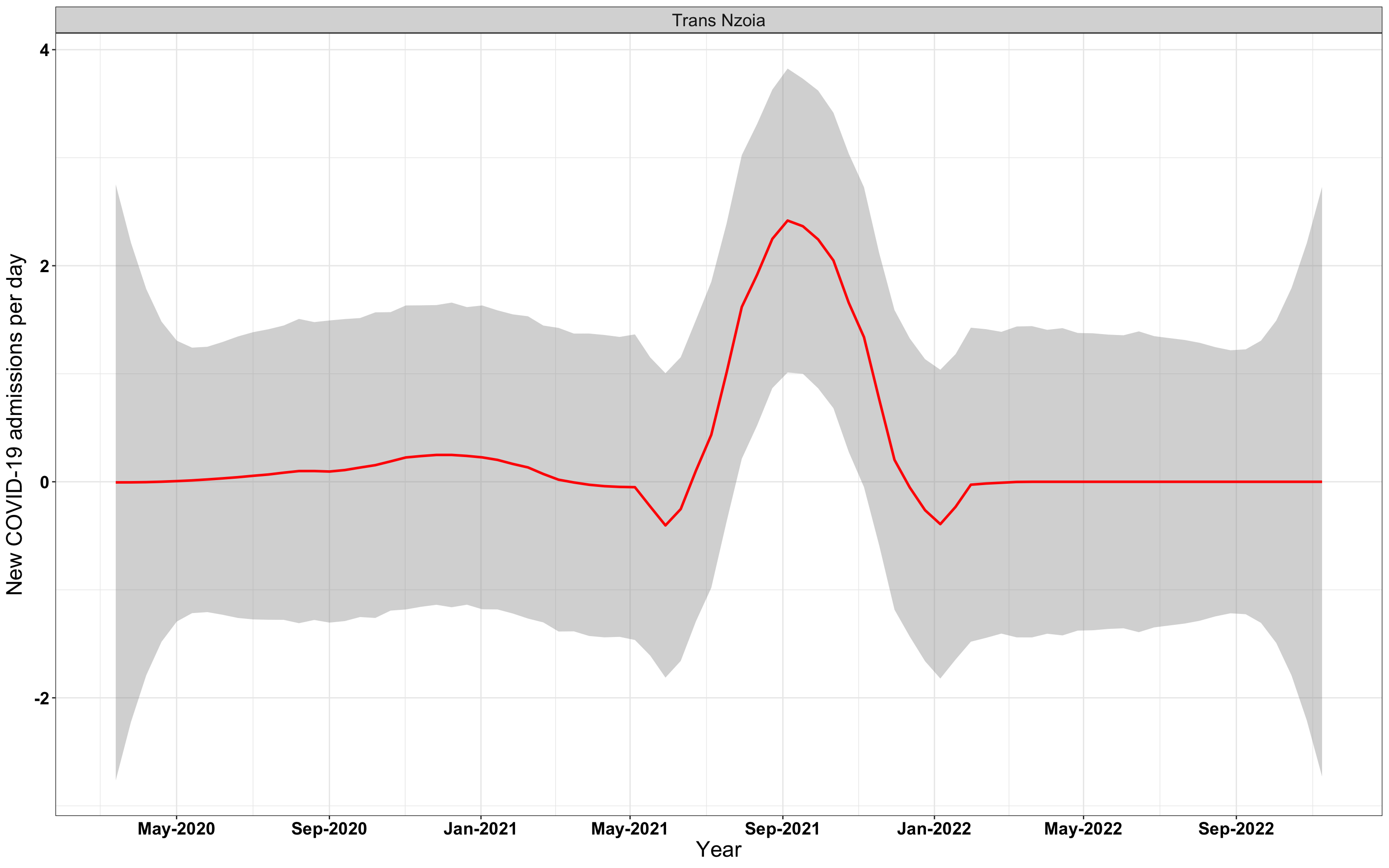
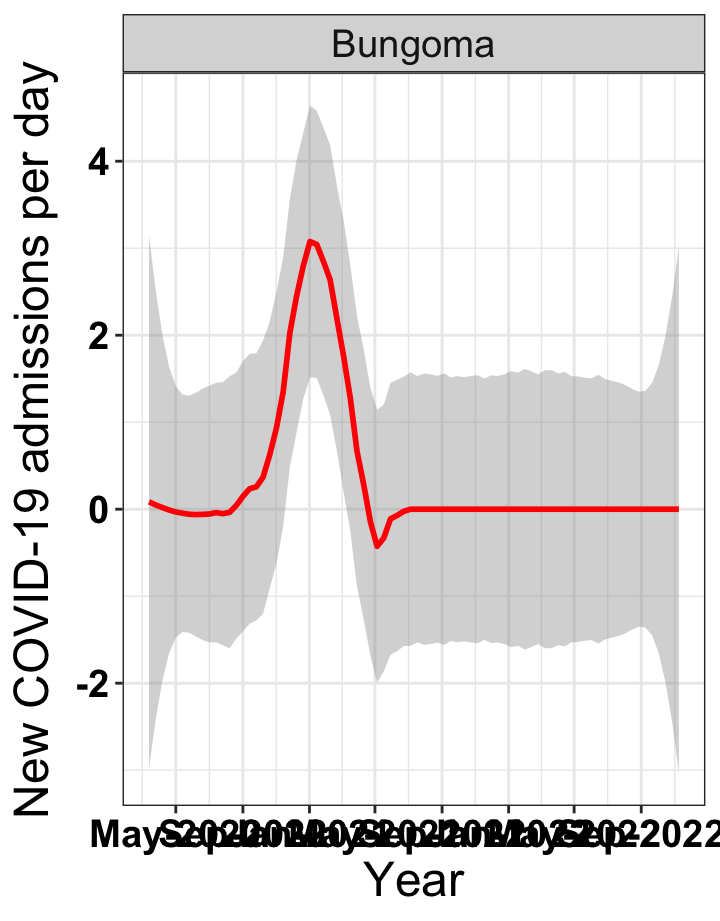
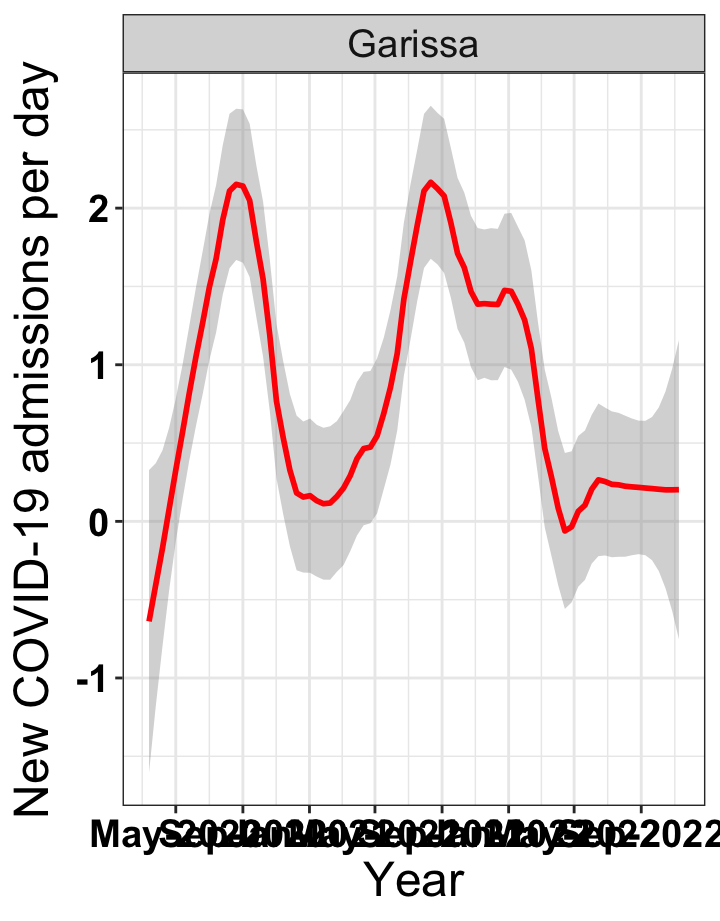
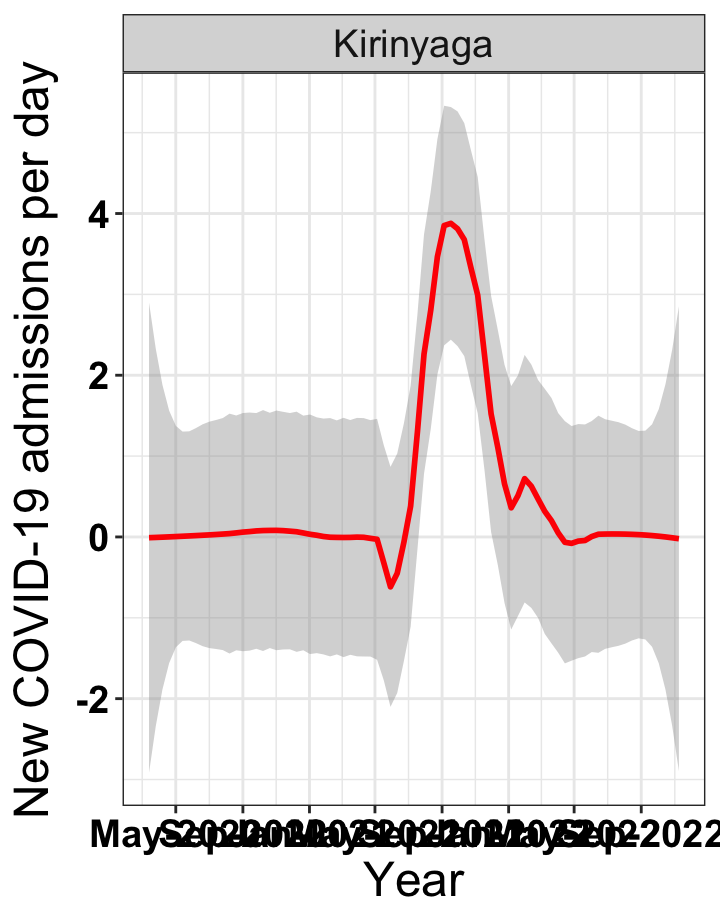
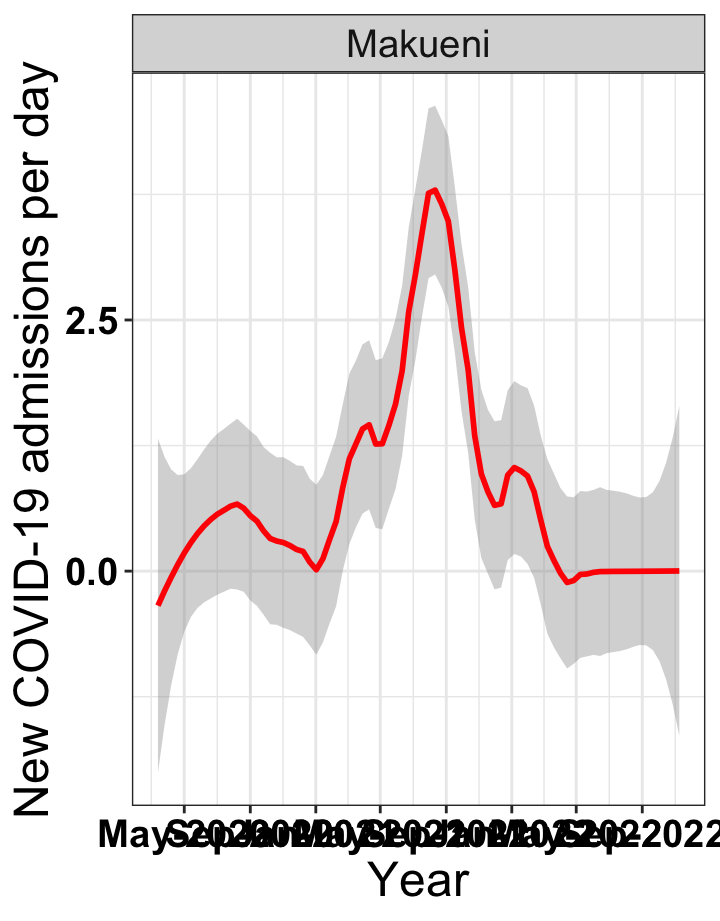
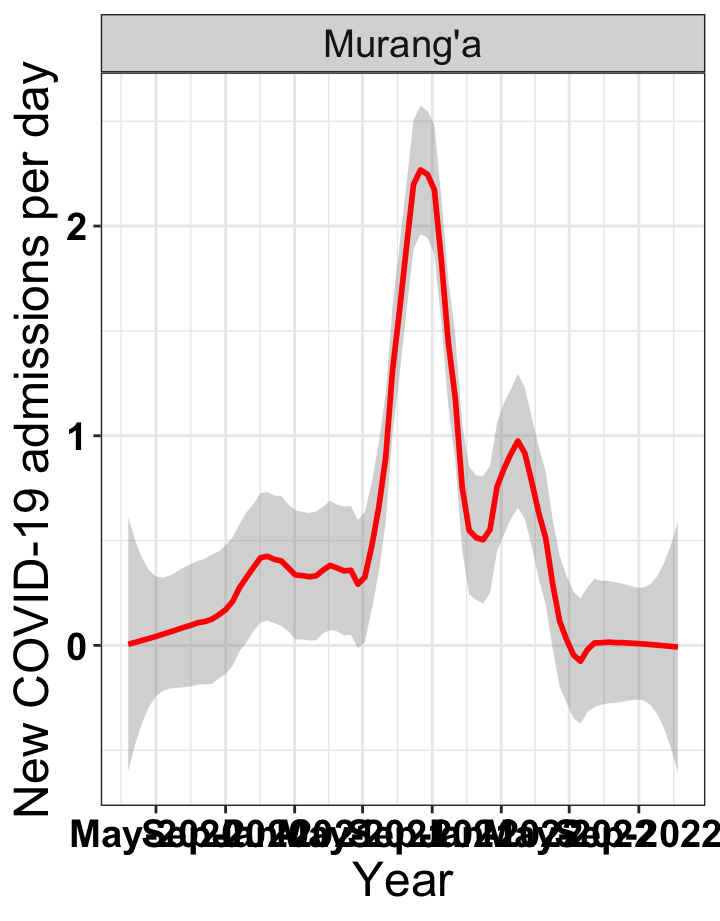
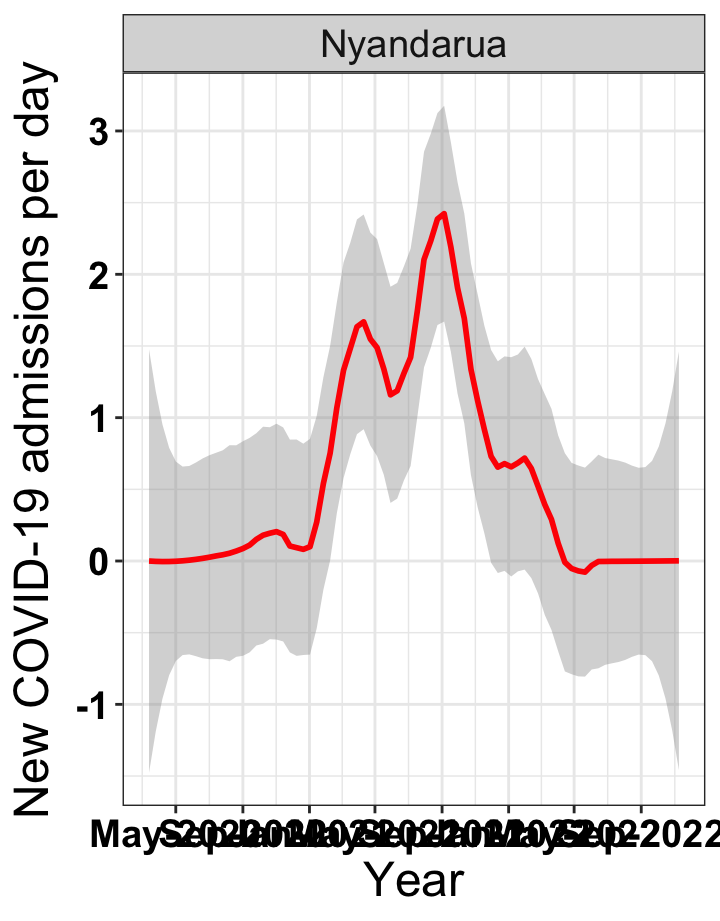
# `r x[2]` {data-navmenu="Hospital admissions"}
```{r,fig.width=16, fig.height=10}
graphs[[2]]
```
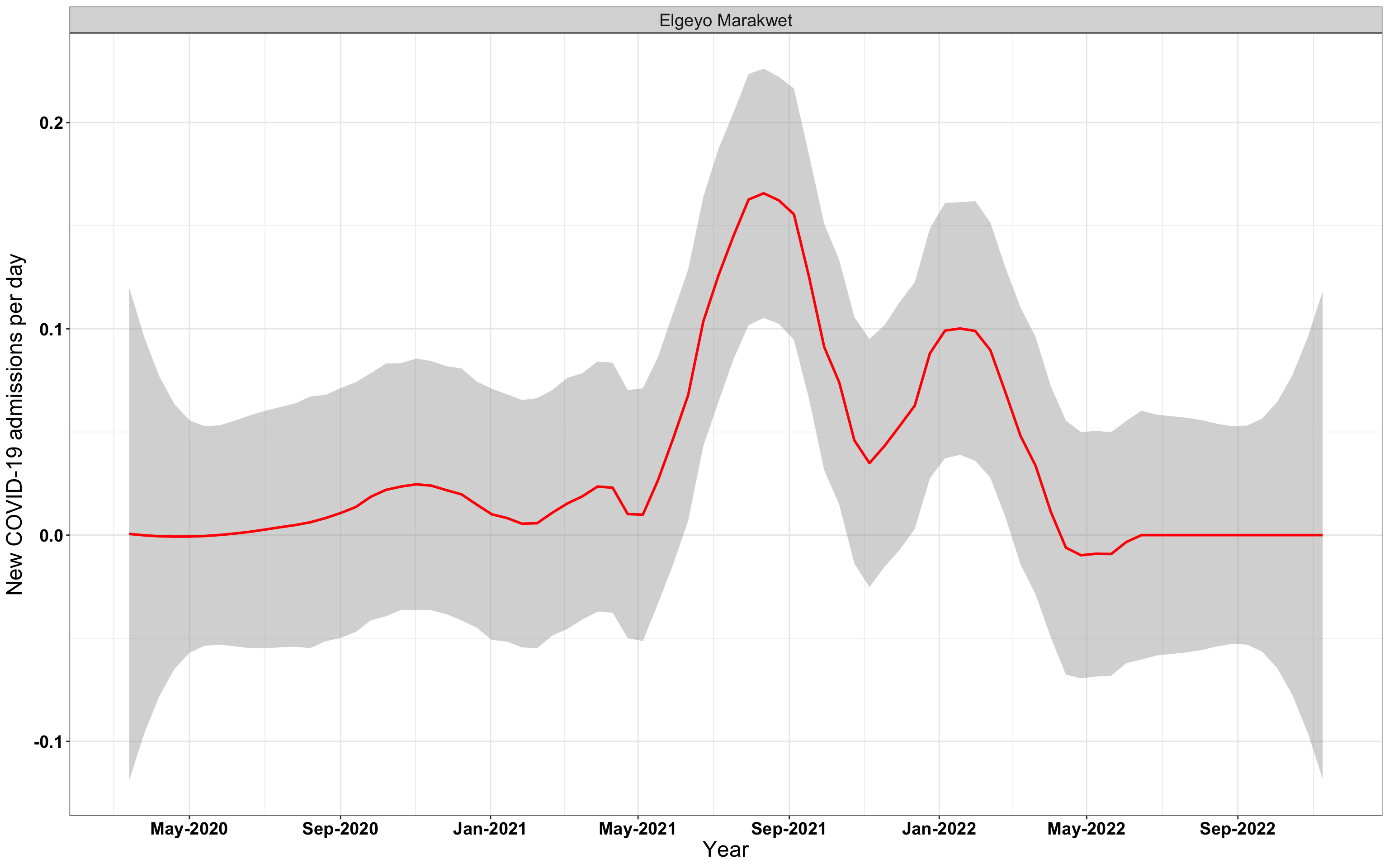
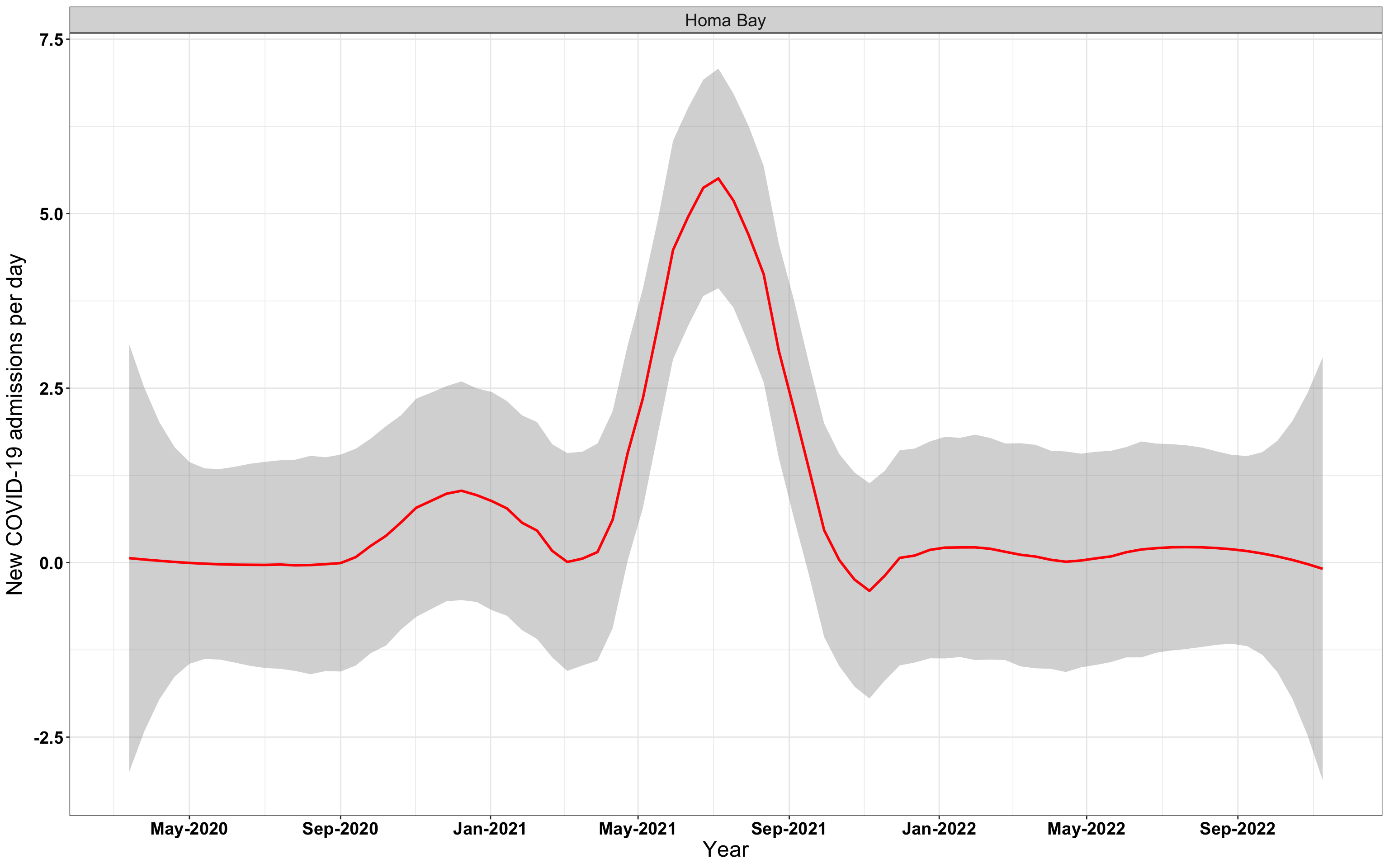
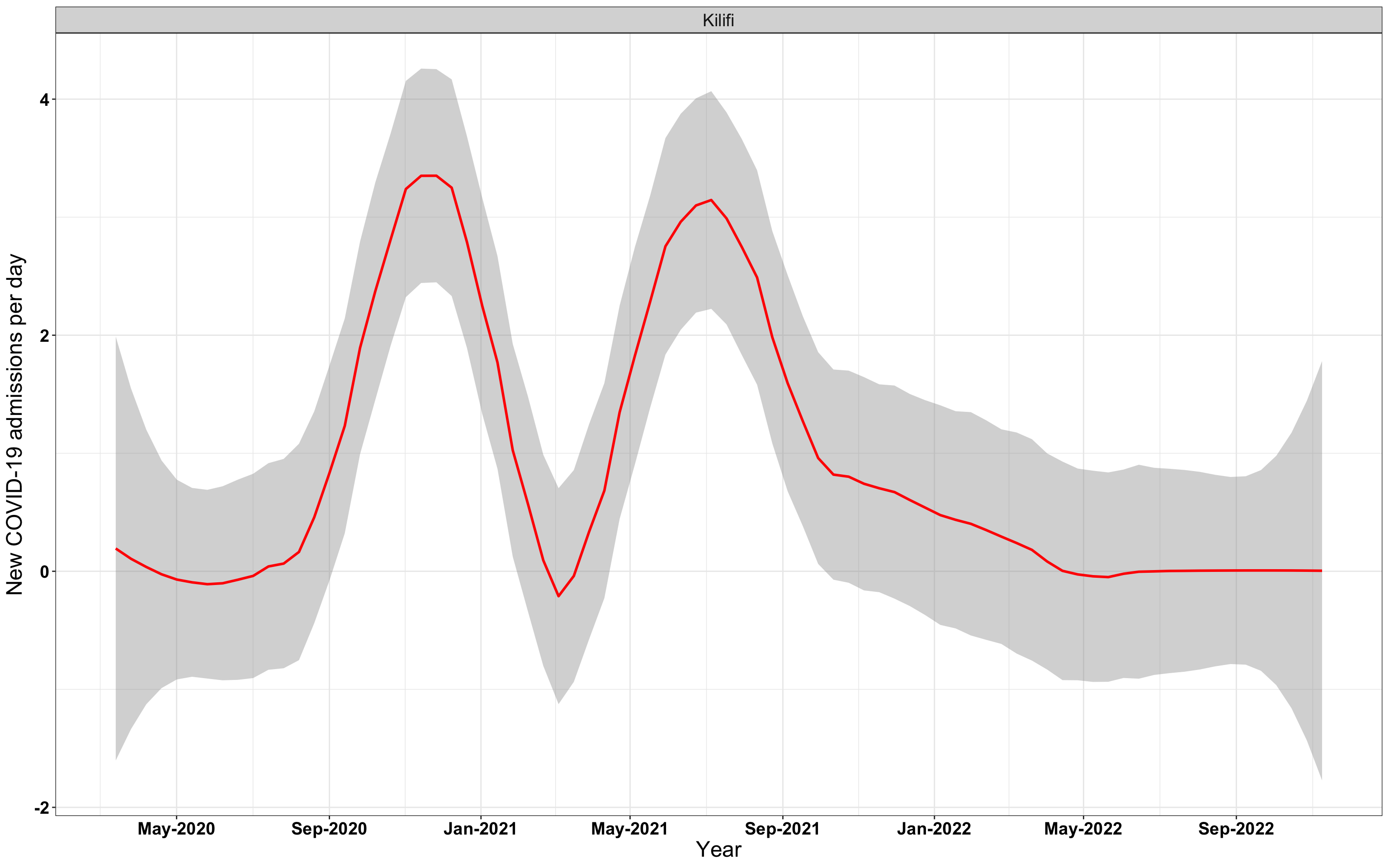
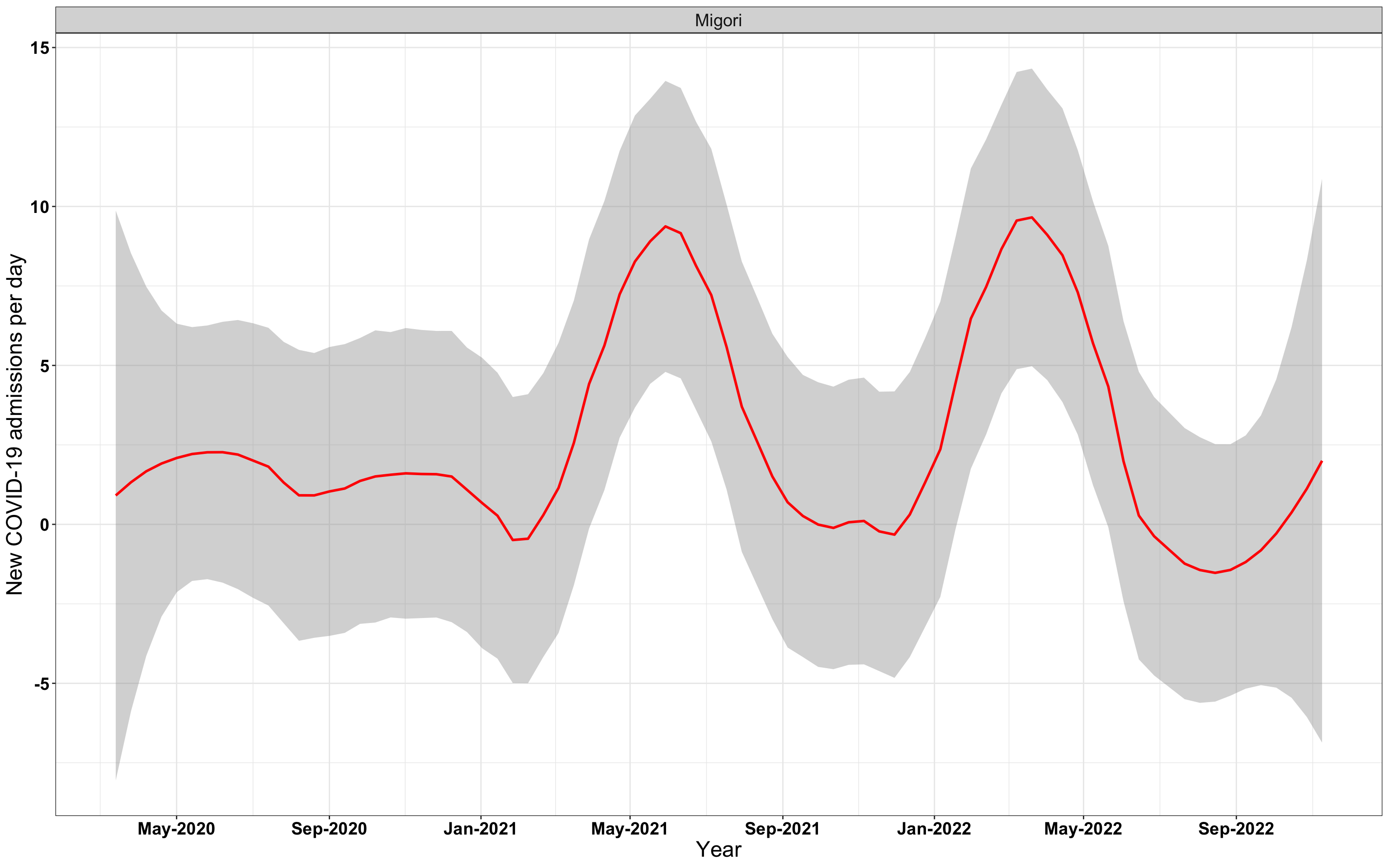
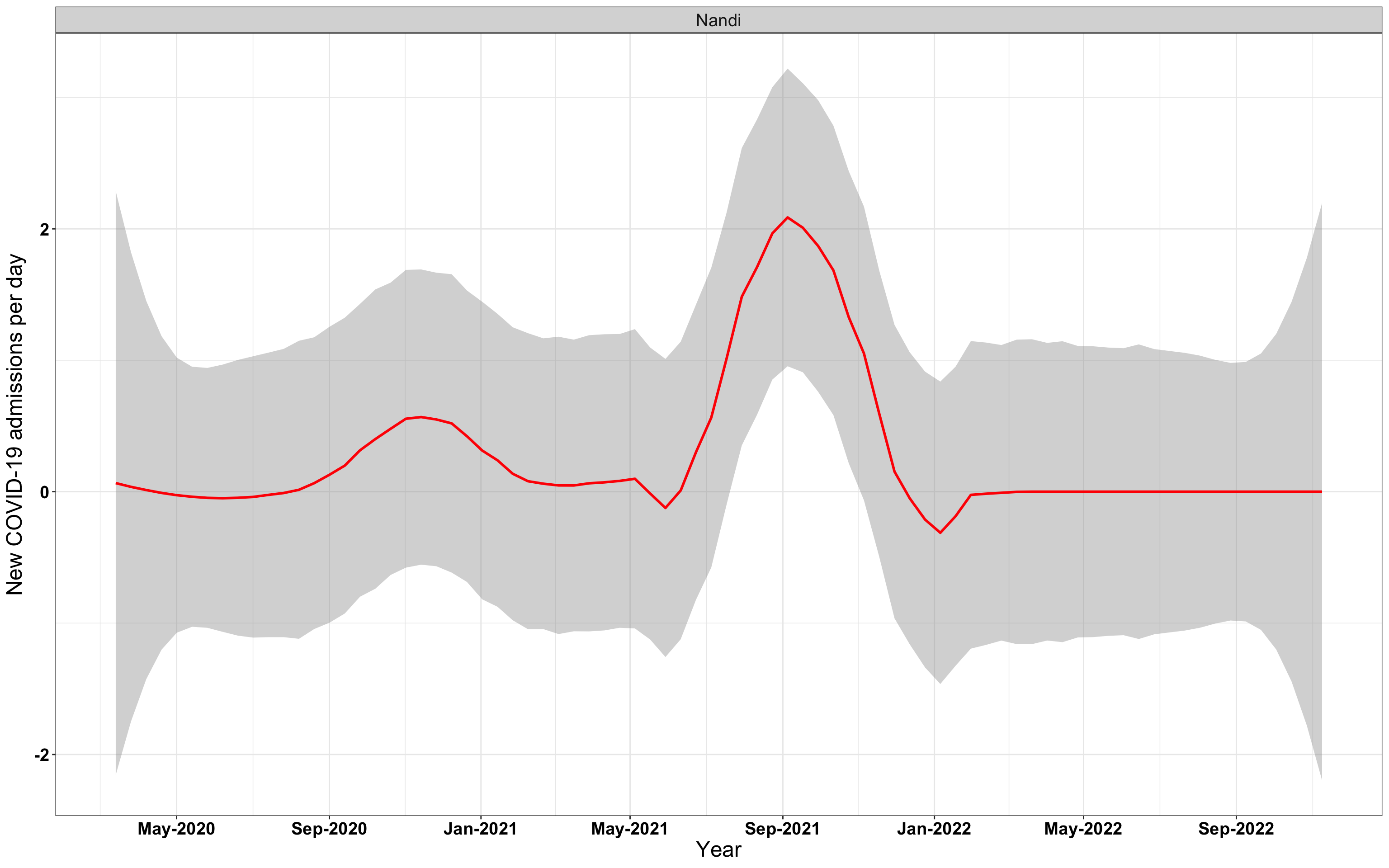
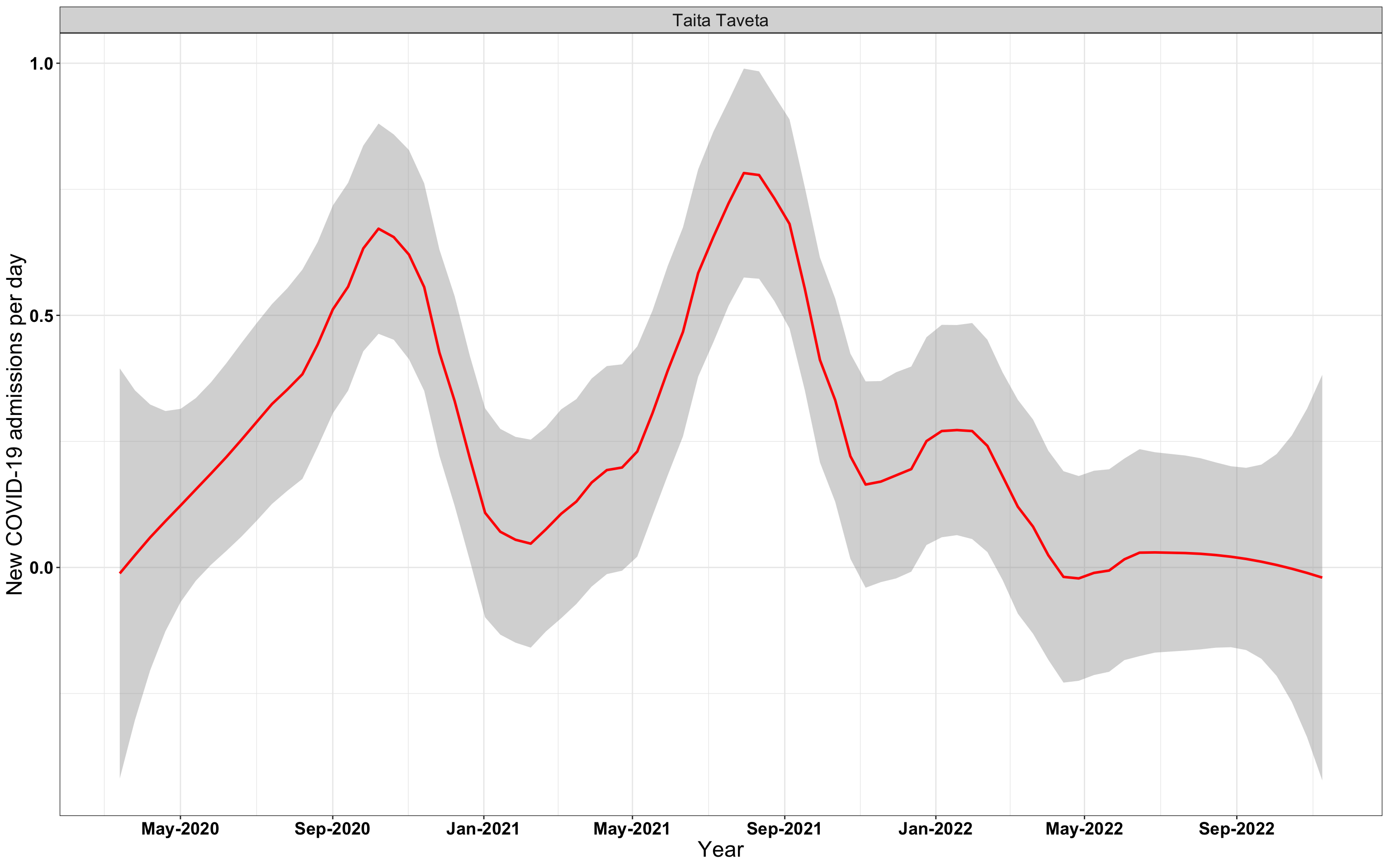
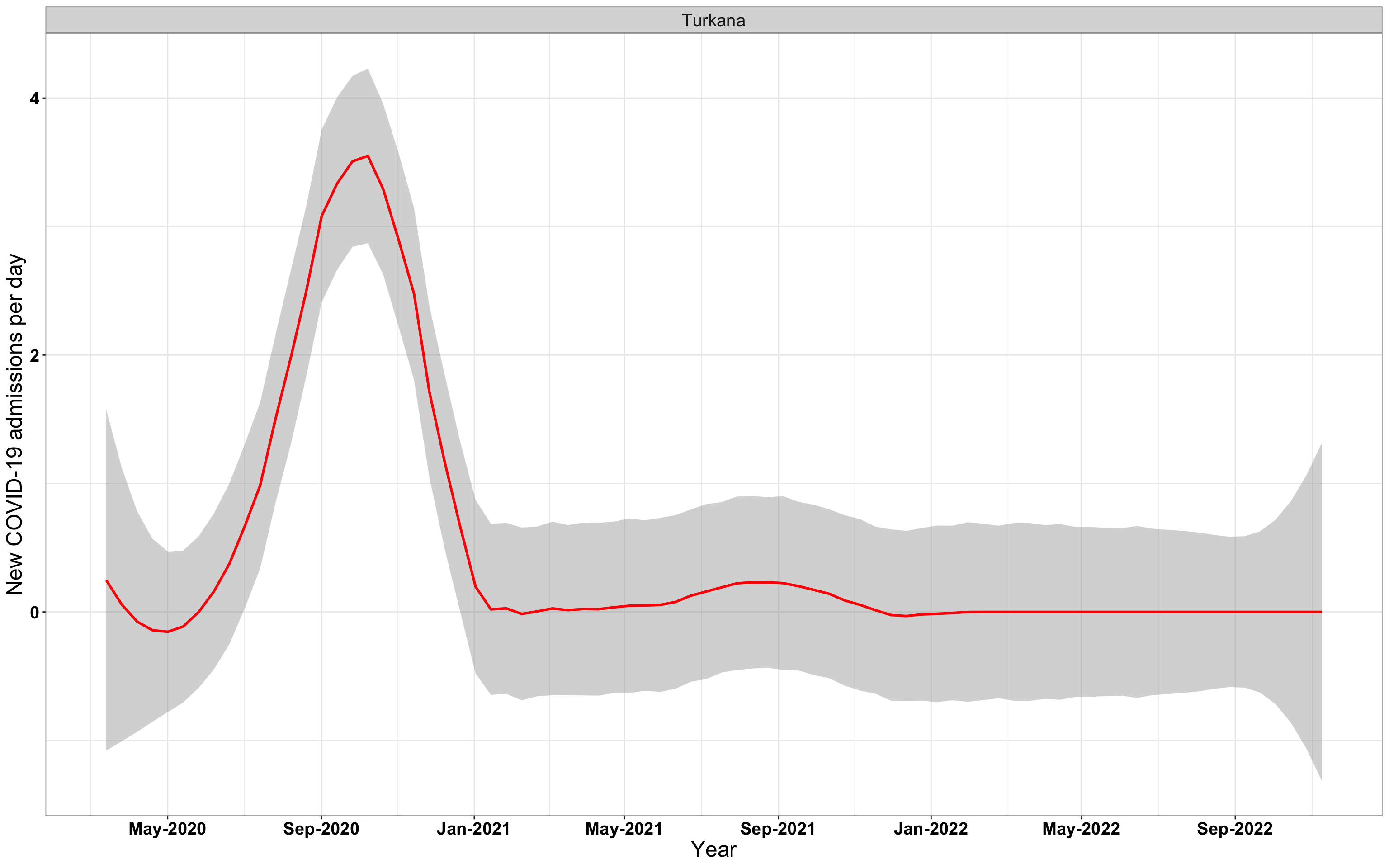
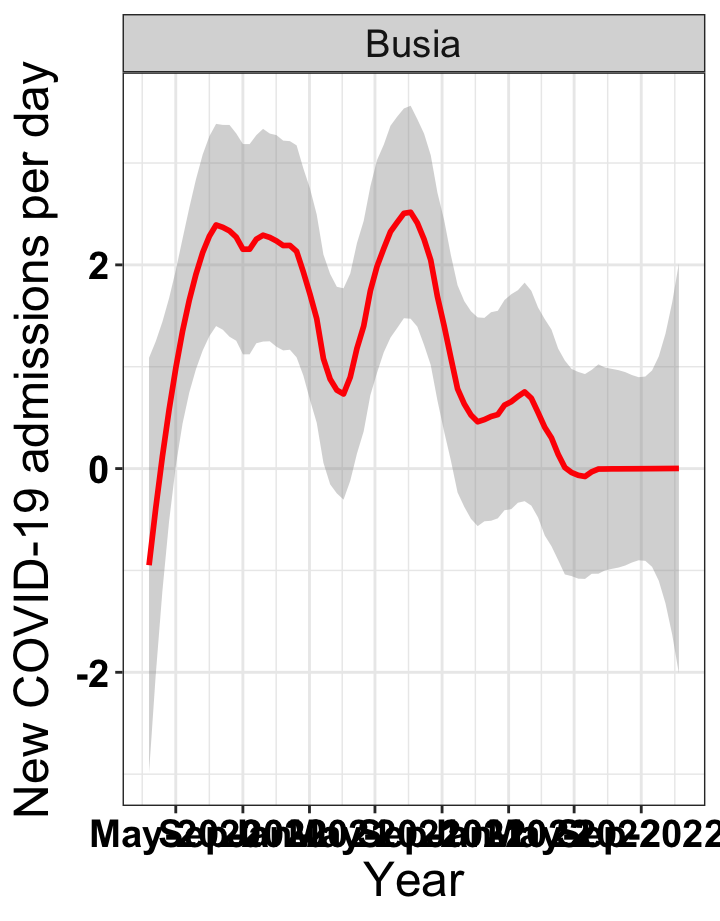
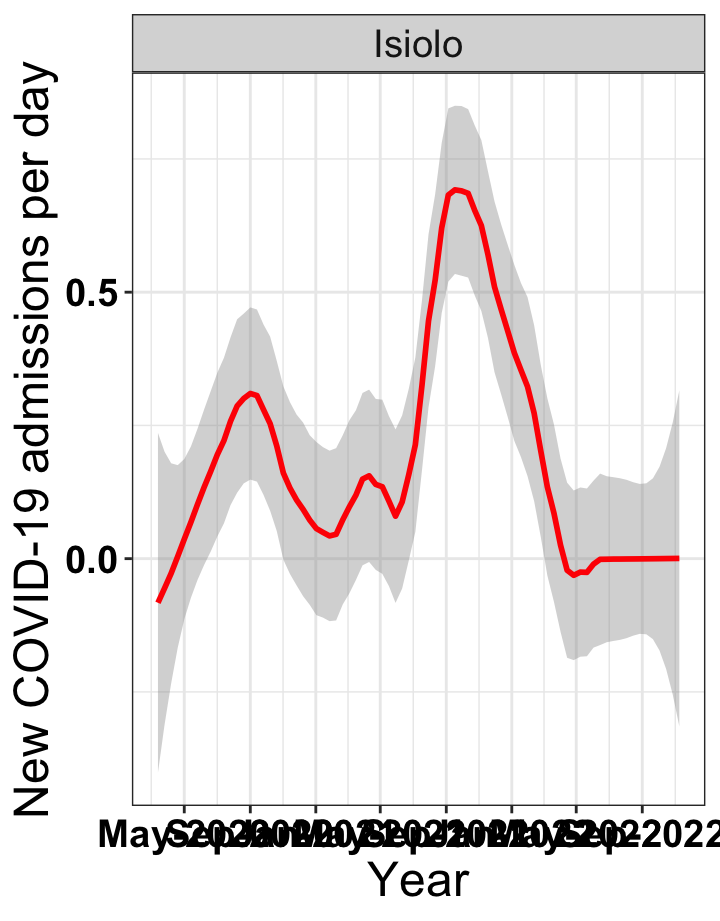
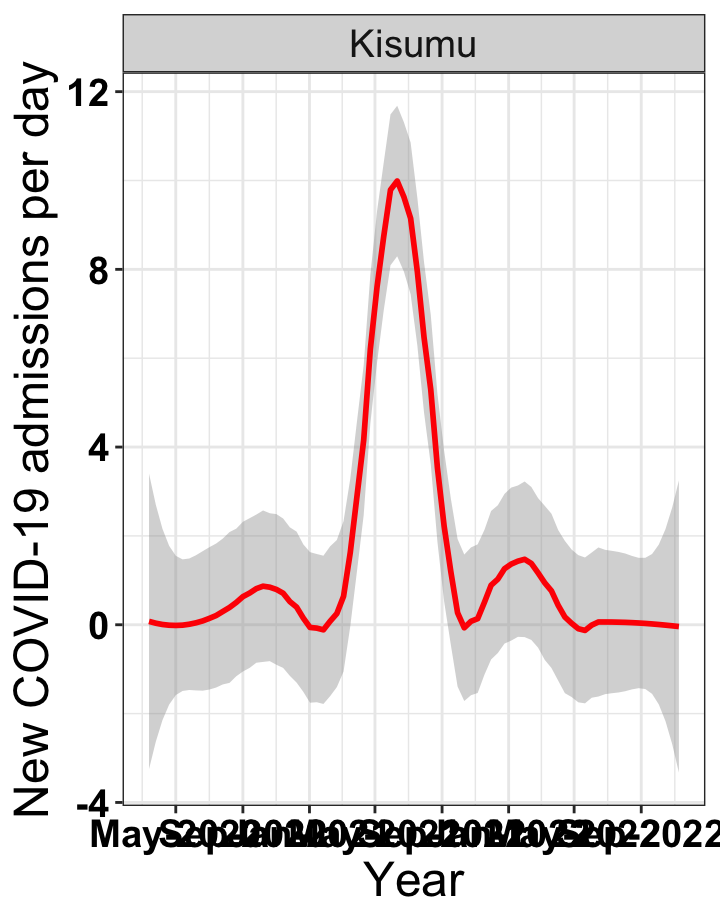
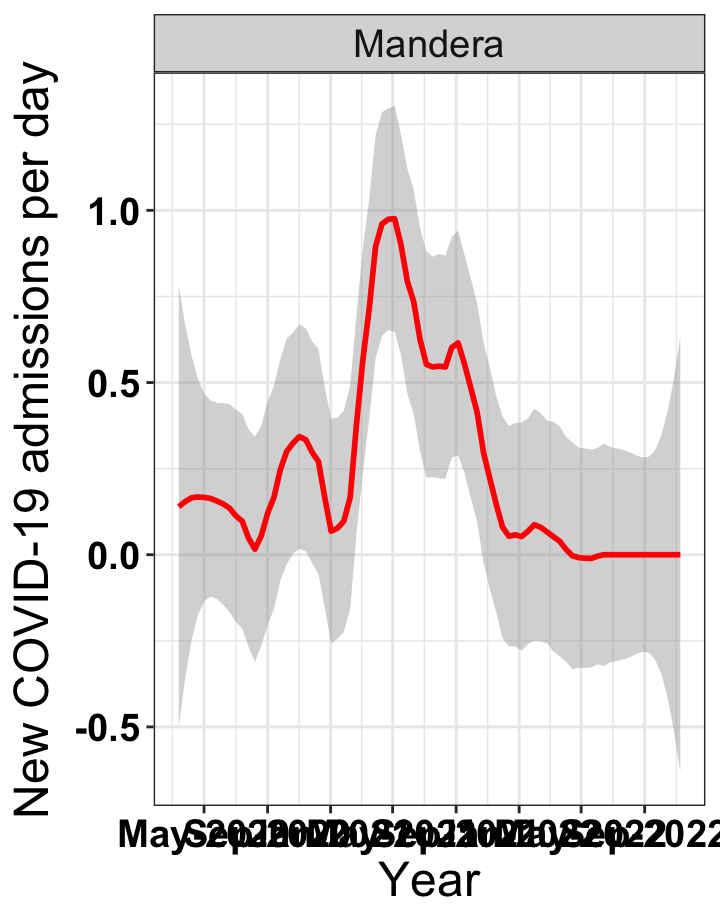
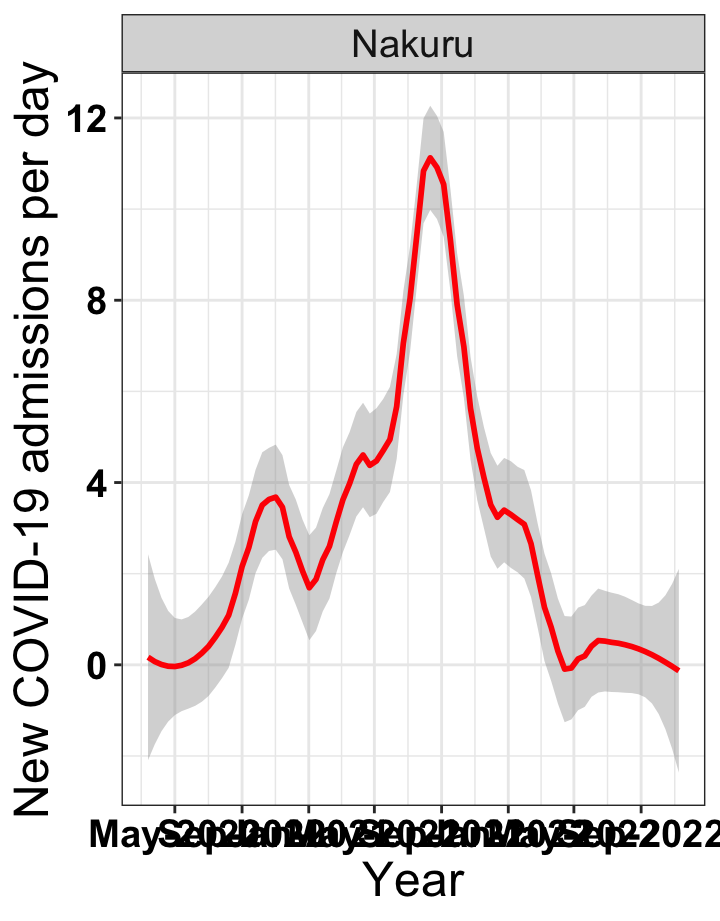
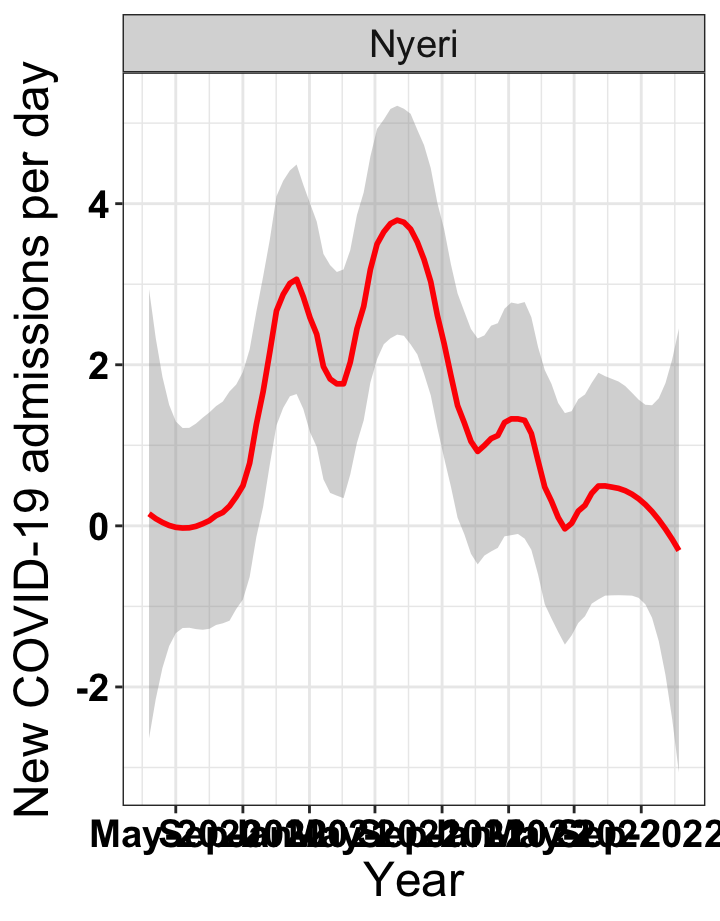
# `r x[3]` {data-navmenu="Hospital admissions"}
```{r,fig.width=16, fig.height=10}
graphs[[3]]
```
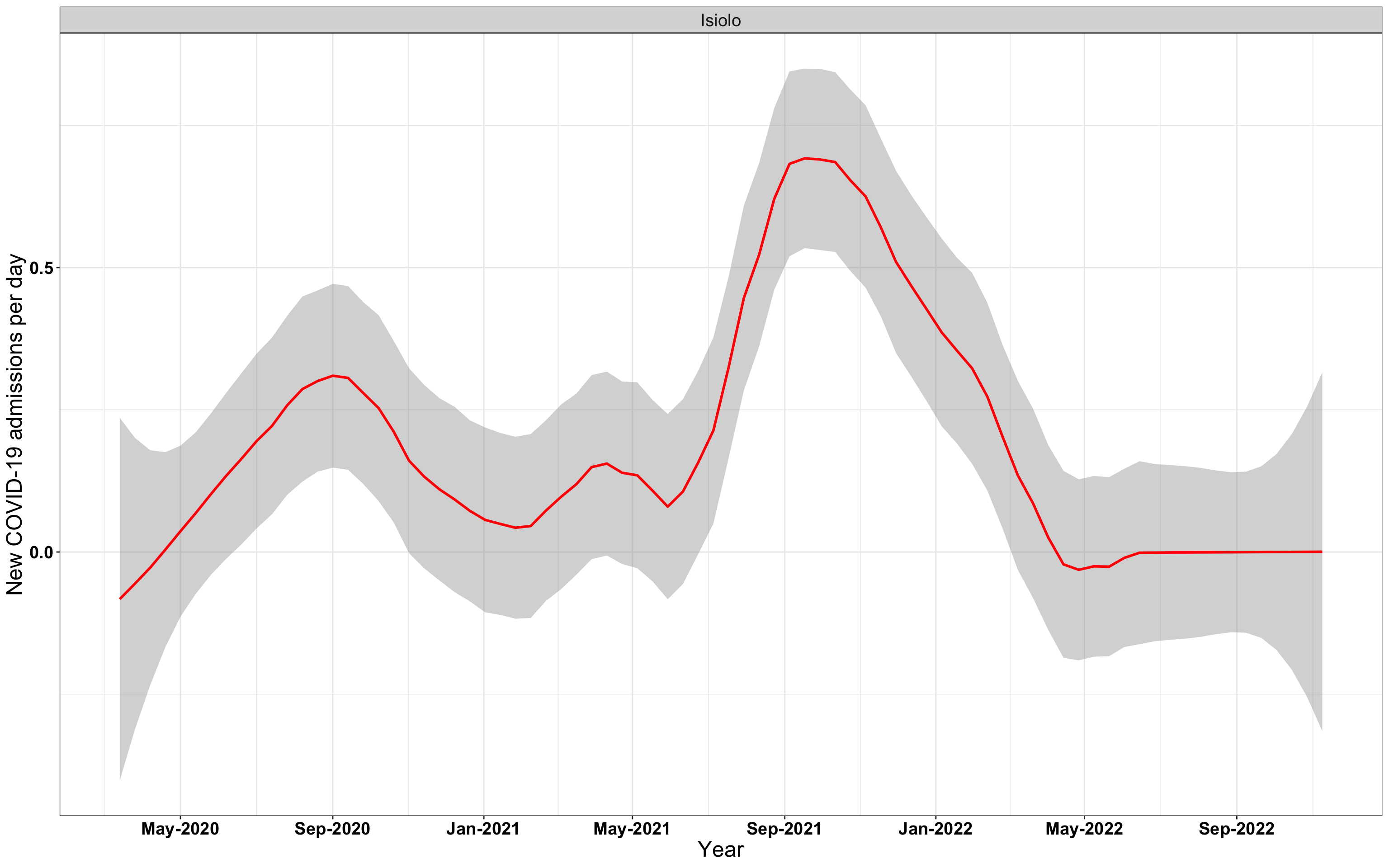
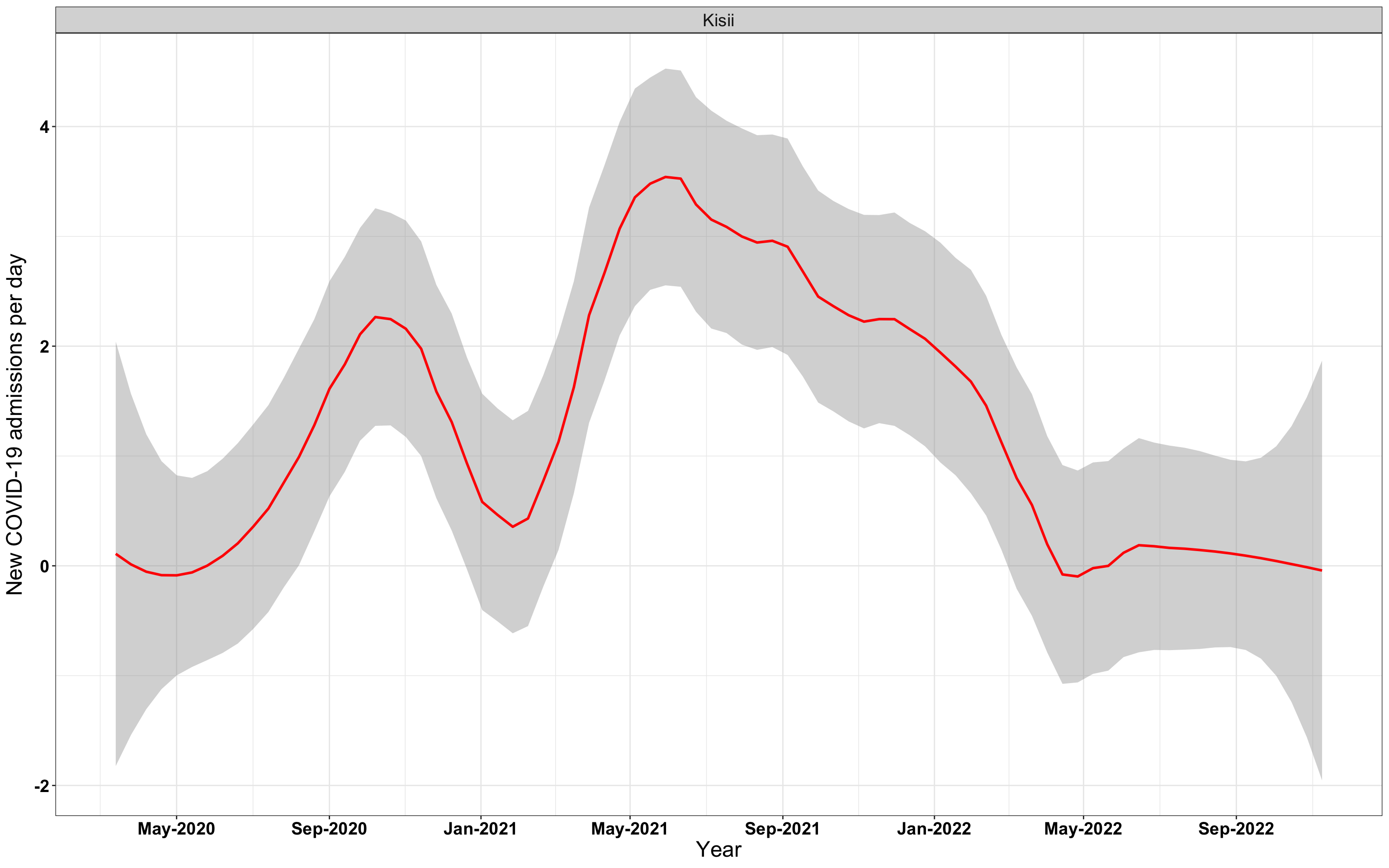
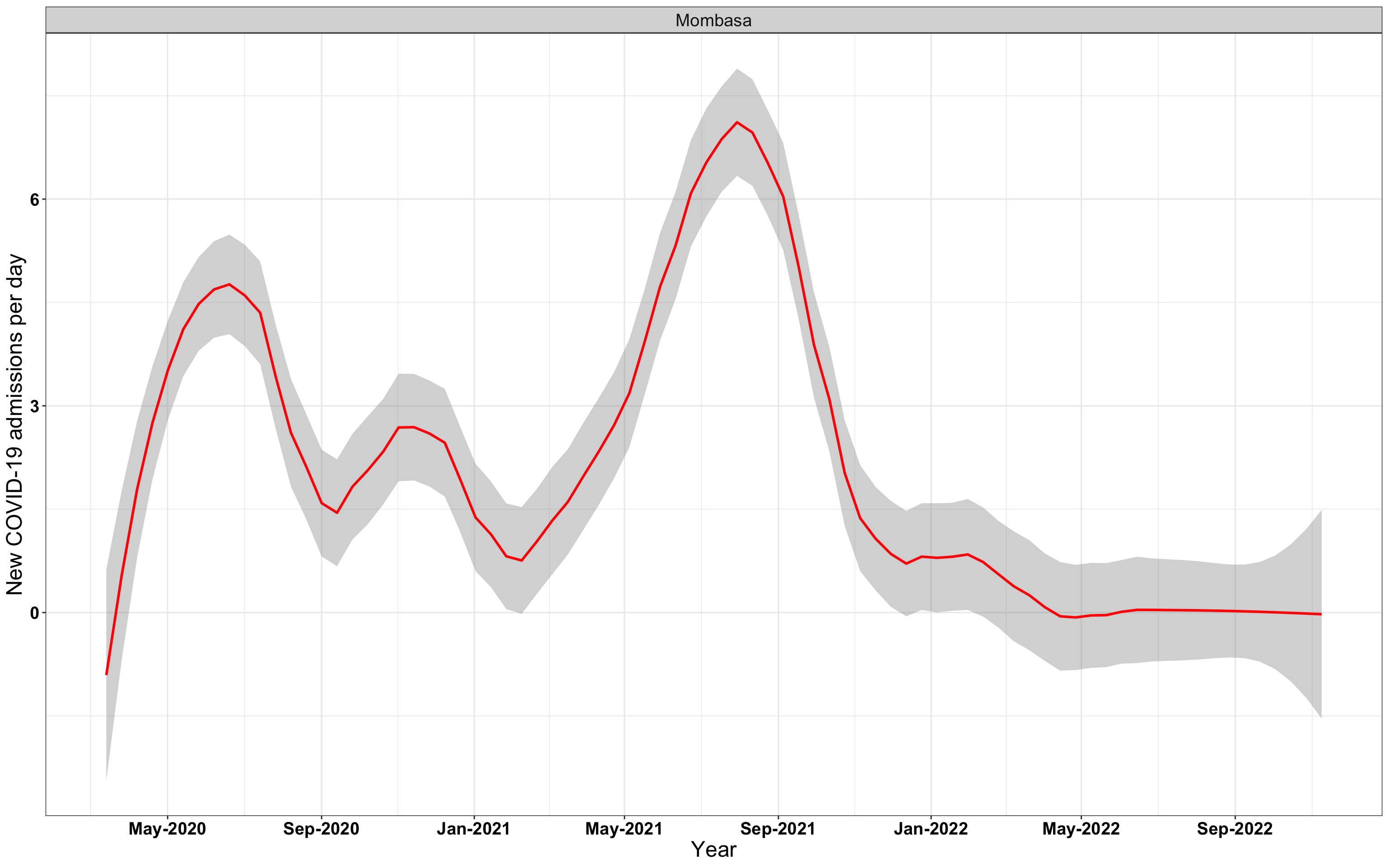
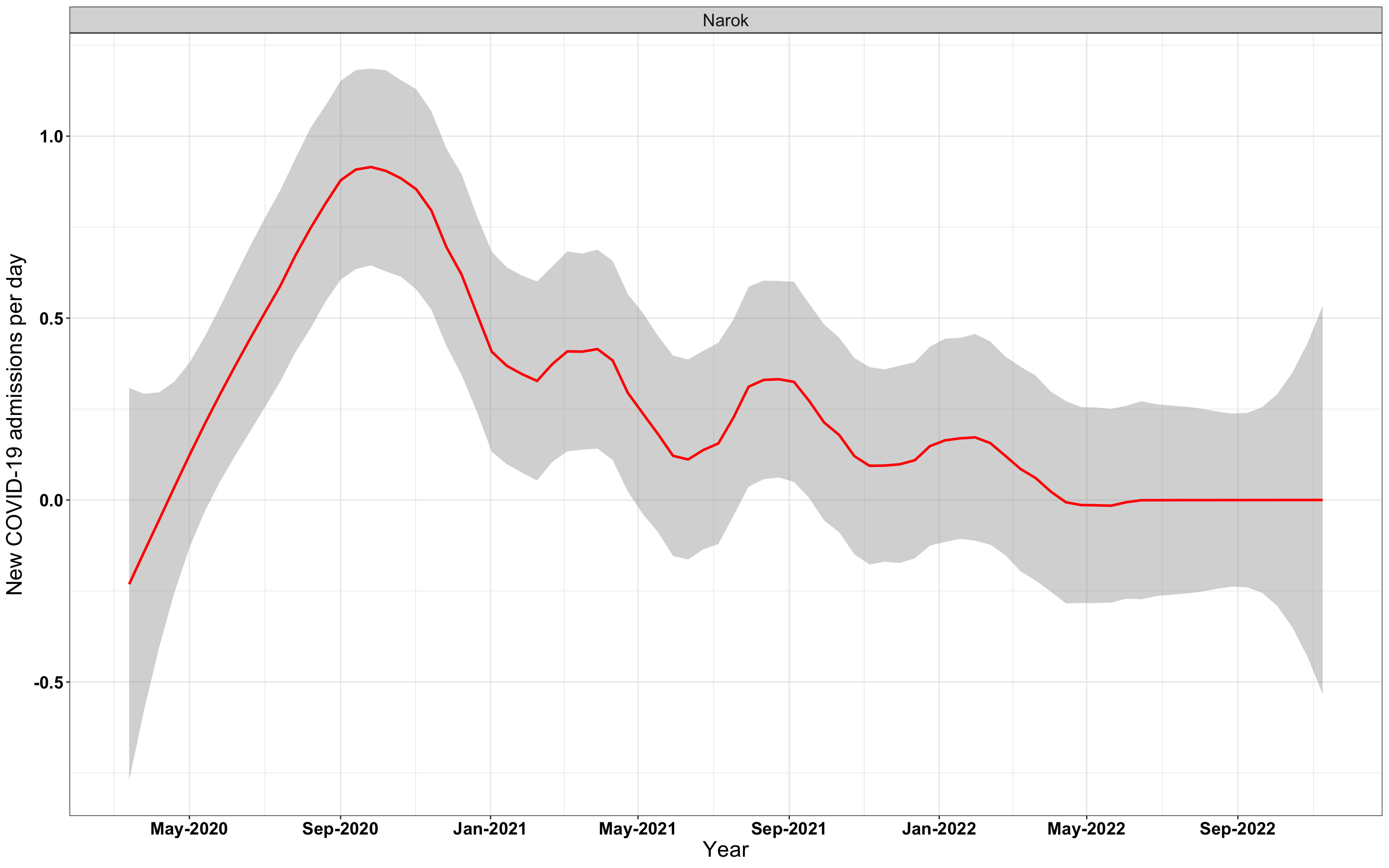
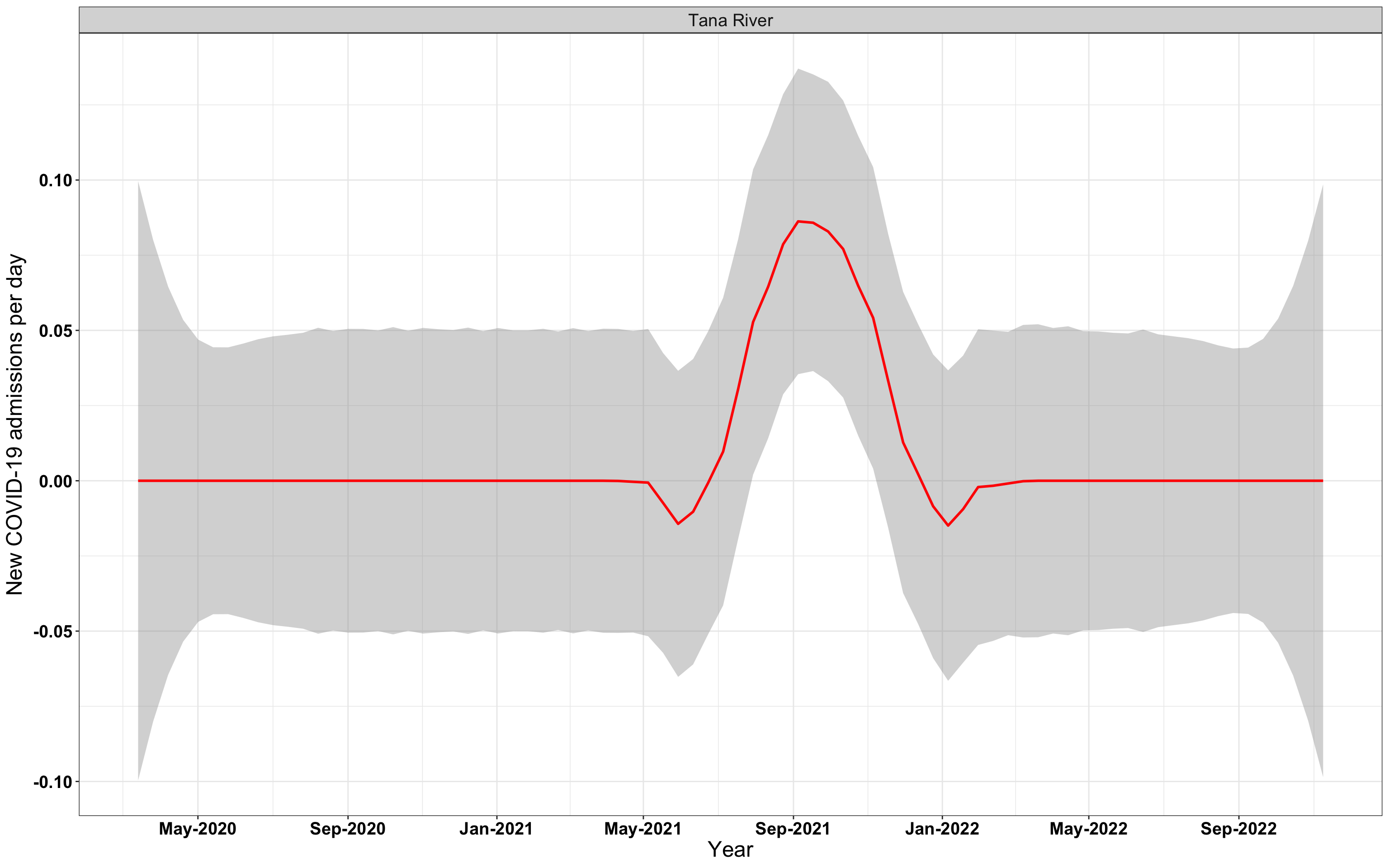
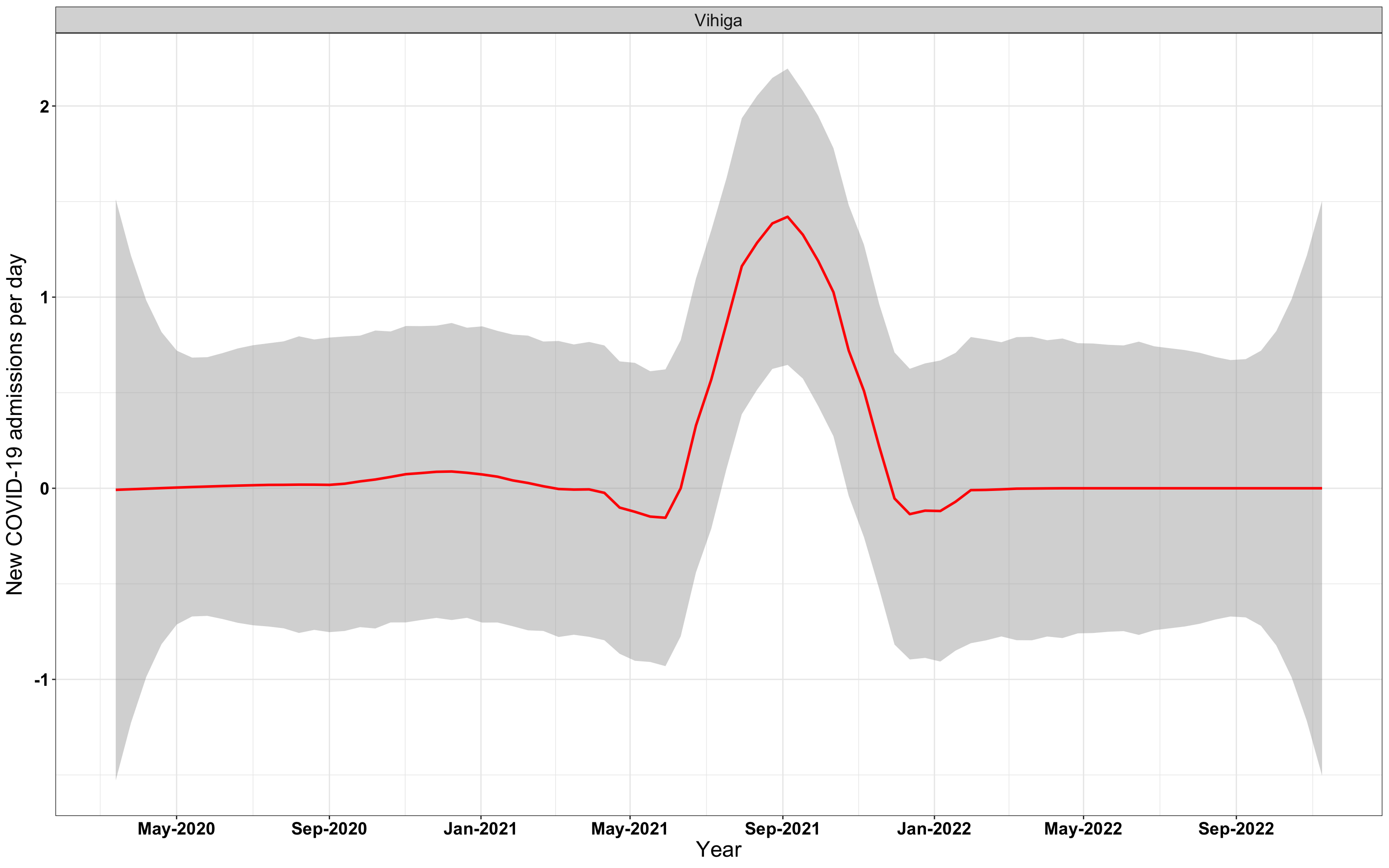
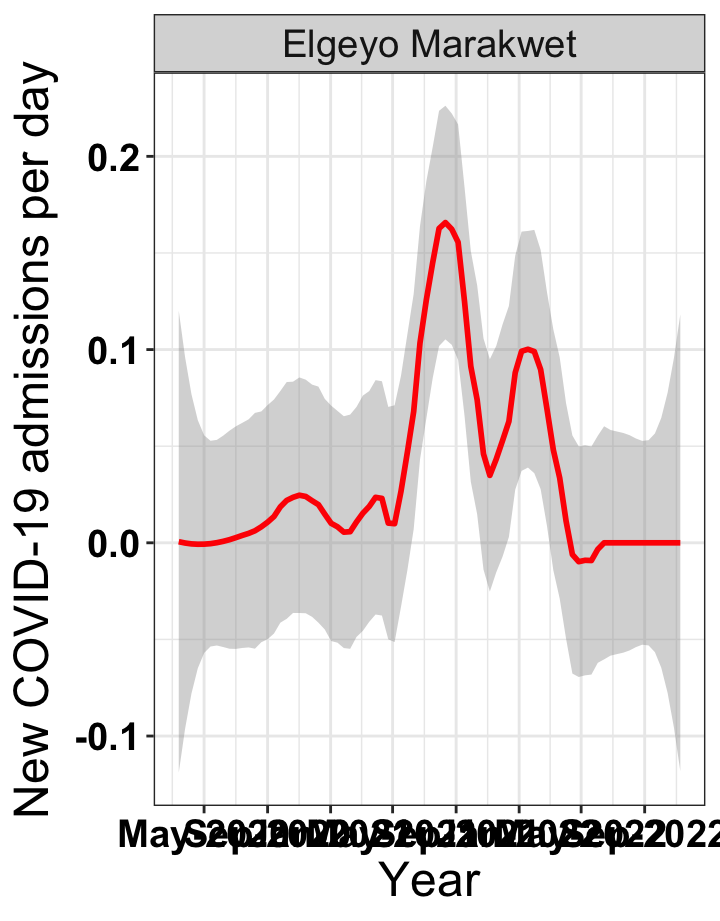
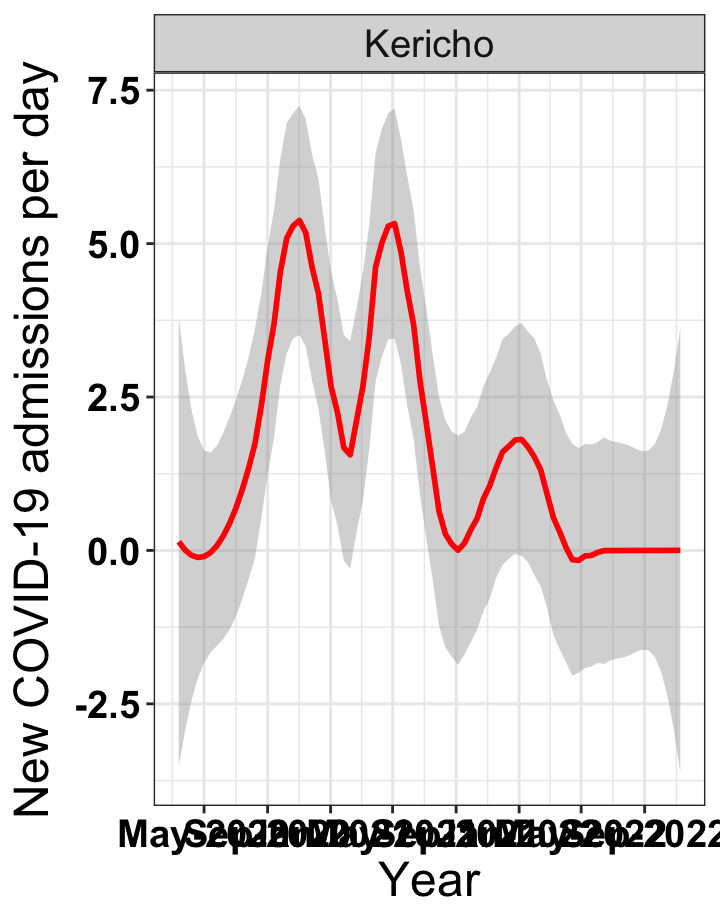
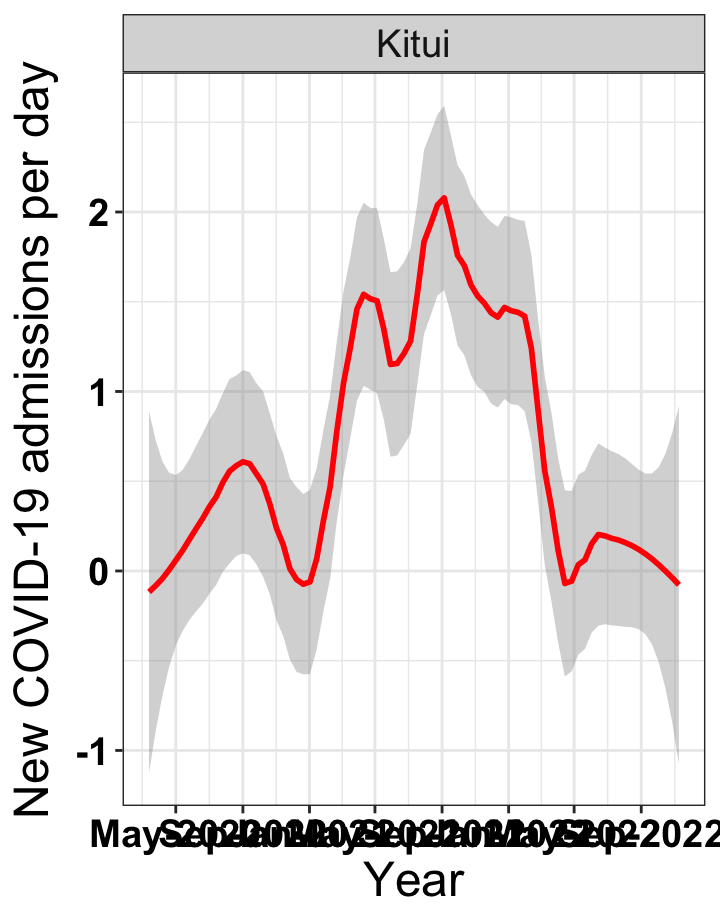
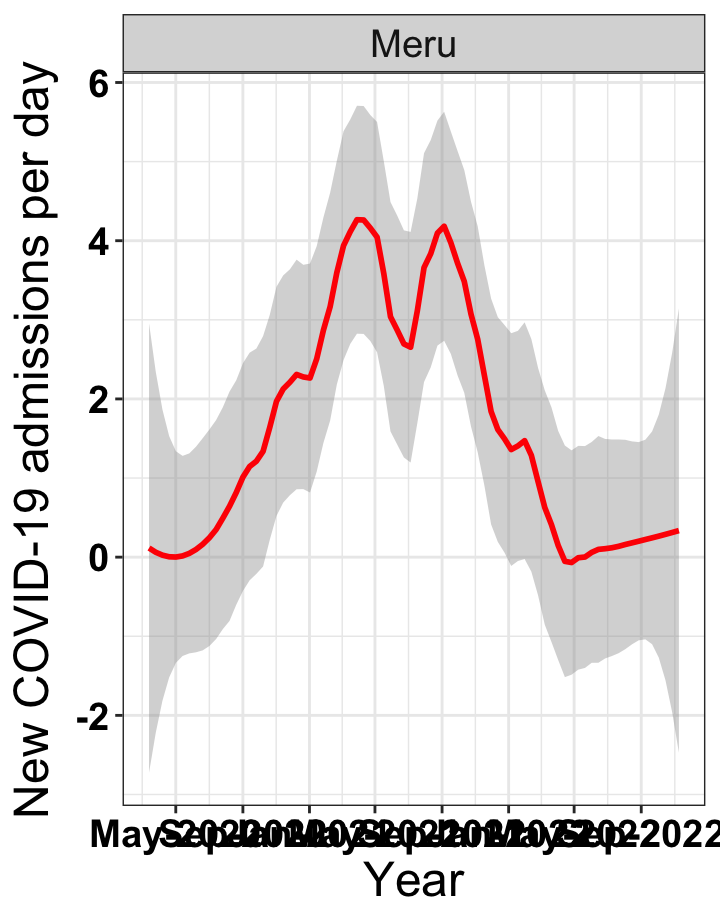
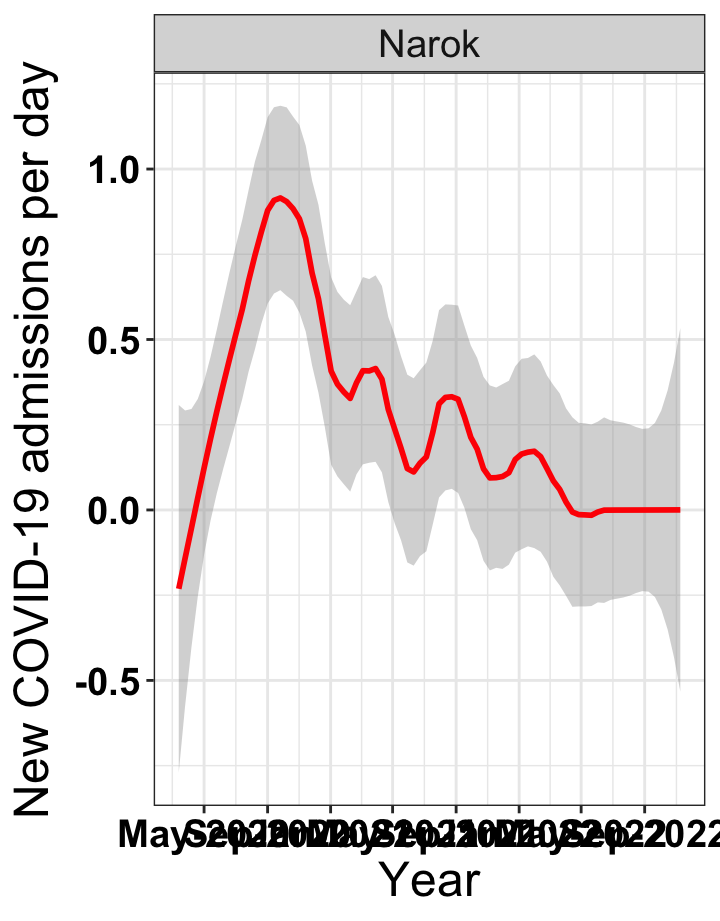
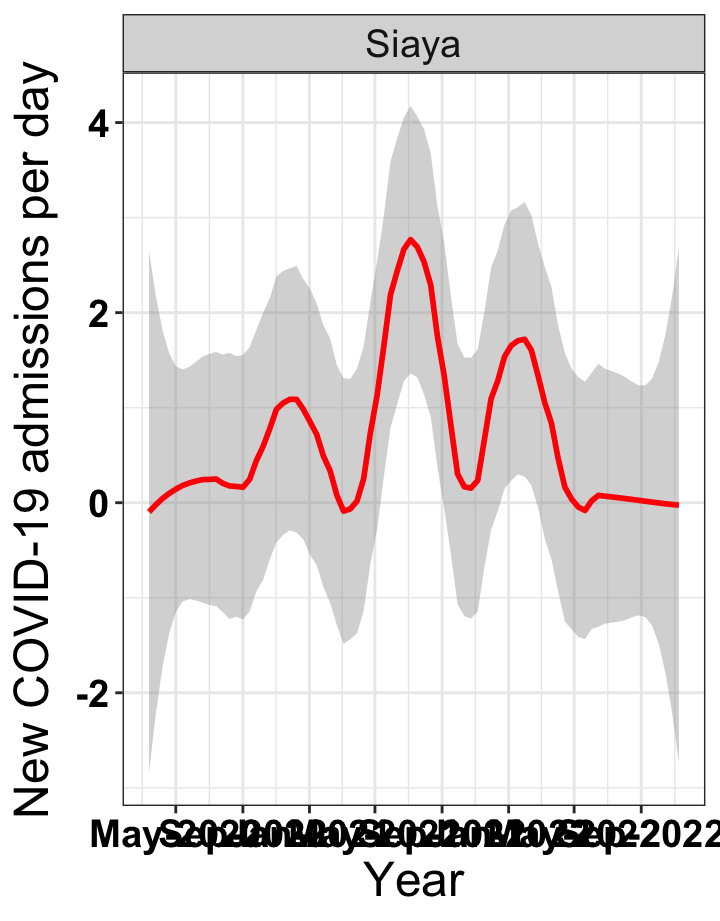
# `r x[4]` {data-navmenu="Hospital admissions"}
```{r,fig.width=16, fig.height=10}
graphs[[4]]
```
# `r x[5]` {data-navmenu="Hospital admissions"}
```{r,fig.width=16, fig.height=10}
graphs[[5]]
```
# `r x[6]` {data-navmenu="Hospital admissions"}
```{r,fig.width=16, fig.height=10}
graphs[[6]]
```
# `r x[7]` {data-navmenu="Hospital admissions"}
```{r,fig.width=16, fig.height=10}
graphs[[7]]
```
# `r x[8]` {data-navmenu="Hospital admissions"}
```{r,fig.width=16, fig.height=10}
graphs[[8]]
```
# `r x[9]` {data-navmenu="Hospital admissions"}
```{r,fig.width=16, fig.height=10}
graphs[[9]]
```
# `r x[10]` {data-navmenu="Hospital admissions"}
```{r,fig.width=16, fig.height=10}
graphs[[10]]
```
# `r x[11]` {data-navmenu="Hospital admissions"}
```{r,fig.width=16, fig.height=10}
graphs[[11]]
```
# `r x[12]` {data-navmenu="Hospital admissions"}
```{r,fig.width=16, fig.height=10}
graphs[[12]]
```
# `r x[13]` {data-navmenu="Hospital admissions"}
```{r,fig.width=16, fig.height=10}
graphs[[13]]
```
# `r x[14]` {data-navmenu="Hospital admissions"}
```{r,fig.width=16, fig.height=10}
graphs[[14]]
```
# `r x[15]` {data-navmenu="Hospital admissions"}
```{r,fig.width=16, fig.height=10}
graphs[[15]]
```
# `r x[16]` {data-navmenu="Hospital admissions"}
```{r,fig.width=16, fig.height=10}
graphs[[16]]
```
# `r x[17]` {data-navmenu="Hospital admissions"}
```{r,fig.width=16, fig.height=10}
graphs[[17]]
```
# `r x[18]` {data-navmenu="Hospital admissions"}
```{r,fig.width=16, fig.height=10}
graphs[[18]]
```
# `r x[19]` {data-navmenu="Hospital admissions"}
```{r,fig.width=16, fig.height=10}
graphs[[19]]
```
# `r x[20]` {data-navmenu="Hospital admissions"}
```{r,fig.width=16, fig.height=10}
graphs[[20]]
```
# `r x[21]` {data-navmenu="Hospital admissions"}
```{r,fig.width=16, fig.height=10}
graphs[[21]]
```
# `r x[22]` {data-navmenu="Hospital admissions"}
```{r,fig.width=16, fig.height=10}
graphs[[22]]
```
# `r x[23]` {data-navmenu="Hospital admissions"}
```{r,fig.width=16, fig.height=10}
graphs[[23]]
```
# `r x[24]` {data-navmenu="Hospital admissions"}
```{r,fig.width=16, fig.height=10}
graphs[[24]]
```
# `r x[25]` {data-navmenu="Hospital admissions"}
```{r,fig.width=16, fig.height=10}
graphs[[25]]
```
# `r x[26]` {data-navmenu="Hospital admissions"}
```{r,fig.width=16, fig.height=10}
graphs[[26]]
```
# `r x[27]` {data-navmenu="Hospital admissions"}
```{r,fig.width=16, fig.height=10}
graphs[[27]]
```
# `r x[28]` {data-navmenu="Hospital admissions"}
```{r,fig.width=16, fig.height=10}
graphs[[28]]
```
# `r x[29]` {data-navmenu="Hospital admissions"}
```{r,fig.width=16, fig.height=10}
graphs[[29]]
```
# `r x[30]` {data-navmenu="Hospital admissions"}
```{r,fig.width=16, fig.height=10}
graphs[[30]]
```
# `r x[31]` {data-navmenu="Hospital admissions"}
```{r,fig.width=16, fig.height=10}
graphs[[31]]
```
# `r x[32]` {data-navmenu="Hospital admissions"}
```{r,fig.width=16, fig.height=10}
graphs[[32]]
```
# `r x[33]` {data-navmenu="Hospital admissions"}
```{r,fig.width=16, fig.height=10}
graphs[[33]]
```
# `r x[34]` {data-navmenu="Hospital admissions"}
```{r,fig.width=16, fig.height=10}
graphs[[34]]
```
# `r x[35]` {data-navmenu="Hospital admissions"}
```{r,fig.width=16, fig.height=10}
graphs[[35]]
```
# `r x[36]` {data-navmenu="Hospital admissions"}
```{r,fig.width=16, fig.height=10}
graphs[[36]]
```
# `r x[37]` {data-navmenu="Hospital admissions"}
```{r,fig.width=16, fig.height=10}
graphs[[37]]
```
# `r x[38]` {data-navmenu="Hospital admissions"}
```{r,fig.width=16, fig.height=10}
graphs[[38]]
```
# `r x[39]` {data-navmenu="Hospital admissions"}
```{r,fig.width=16, fig.height=10}
graphs[[39]]
```
# `r x[40]` {data-navmenu="Hospital admissions"}
```{r,fig.width=16, fig.height=10}
graphs[[40]]
```
# `r x[41]` {data-navmenu="Hospital admissions"}
```{r,fig.width=16, fig.height=10}
graphs[[41]]
```
# `r x[42]` {data-navmenu="Hospital admissions"}
```{r,fig.width=16, fig.height=10}
graphs[[42]]
```
# `r x[43]` {data-navmenu="Hospital admissions"}
```{r,fig.width=16, fig.height=10}
graphs[[43]]
```
# `r x[44]` {data-navmenu="Hospital admissions"}
```{r,fig.width=16, fig.height=10}
graphs[[44]]
```
# `r x[45]` {data-navmenu="Hospital admissions"}
```{r,fig.width=16, fig.height=10}
graphs[[45]]
```
# `r x[46]` {data-navmenu="Hospital admissions"}
```{r,fig.width=16, fig.height=10}
graphs[[46]]
```
# `r x[47]` {data-navmenu="Hospital admissions"}
```{r,fig.width=16, fig.height=10}
graphs[[47]]
```
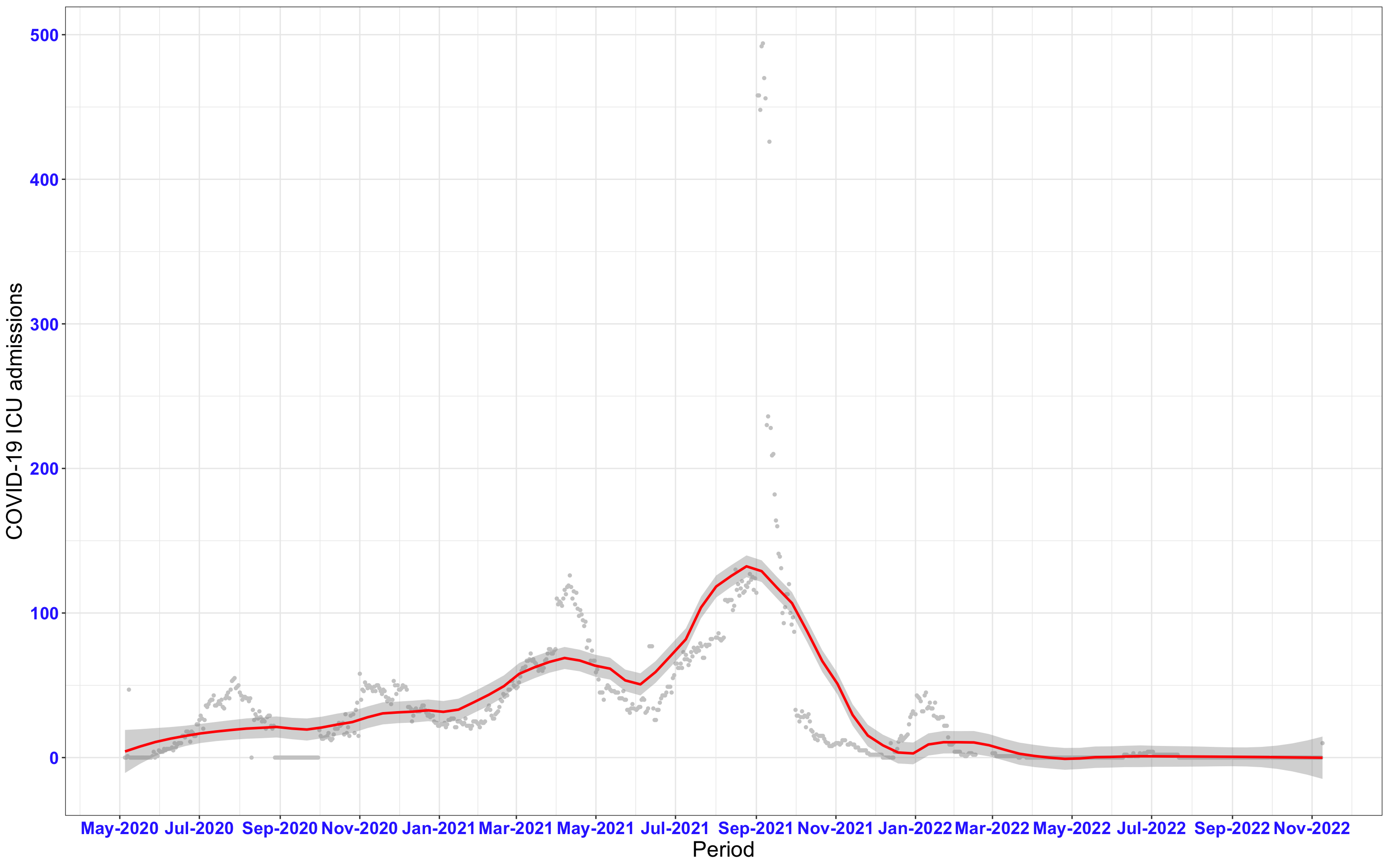
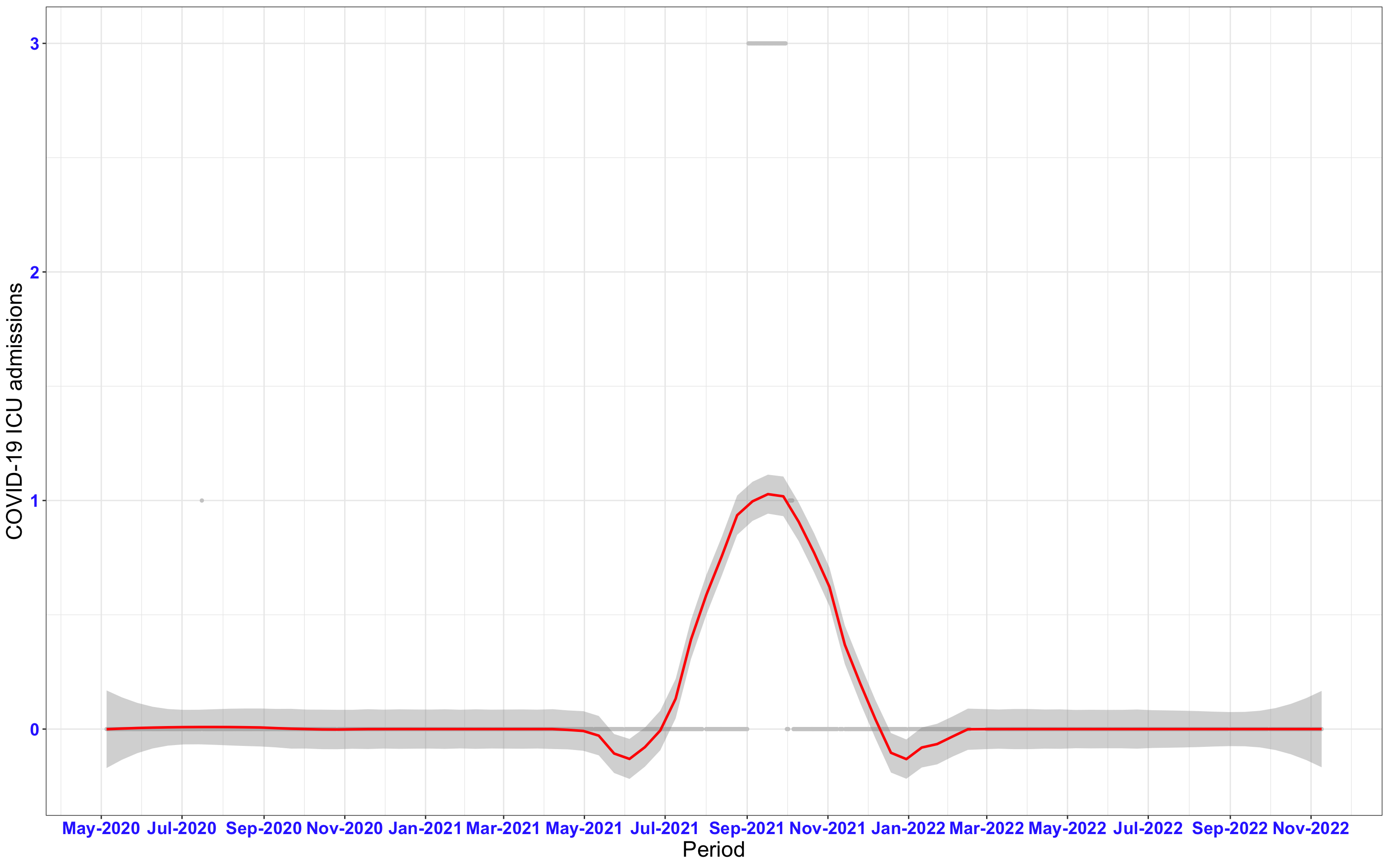
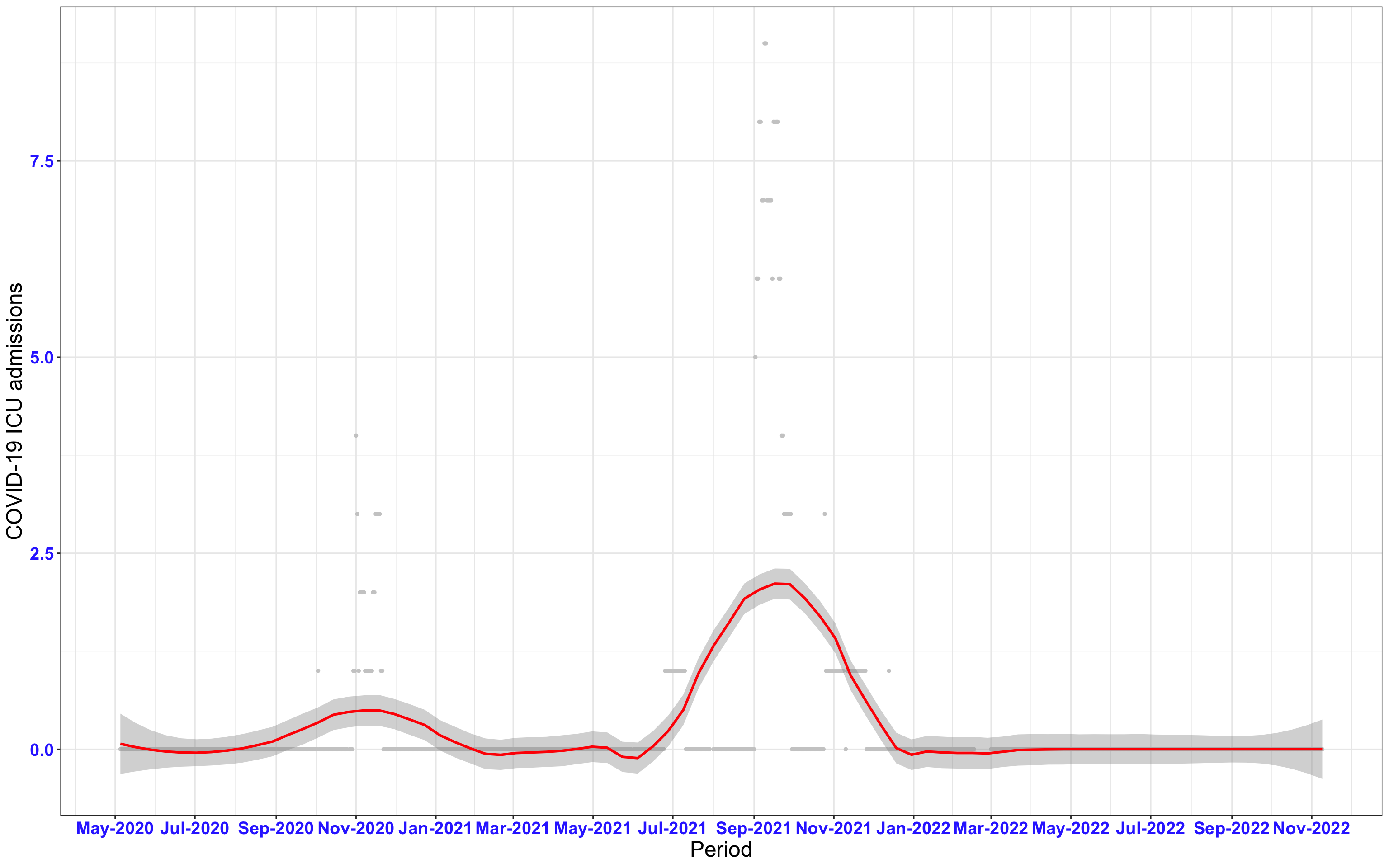
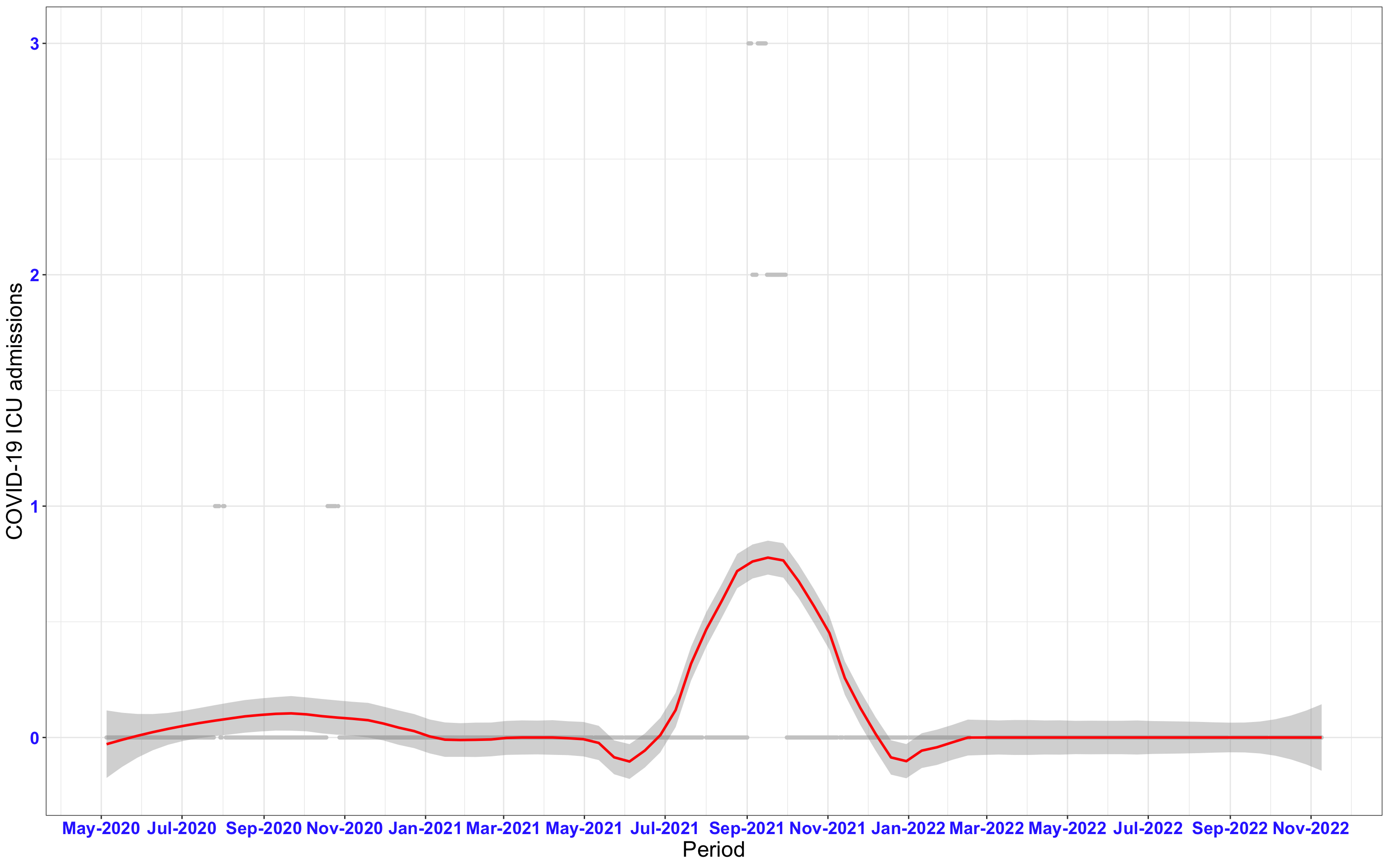
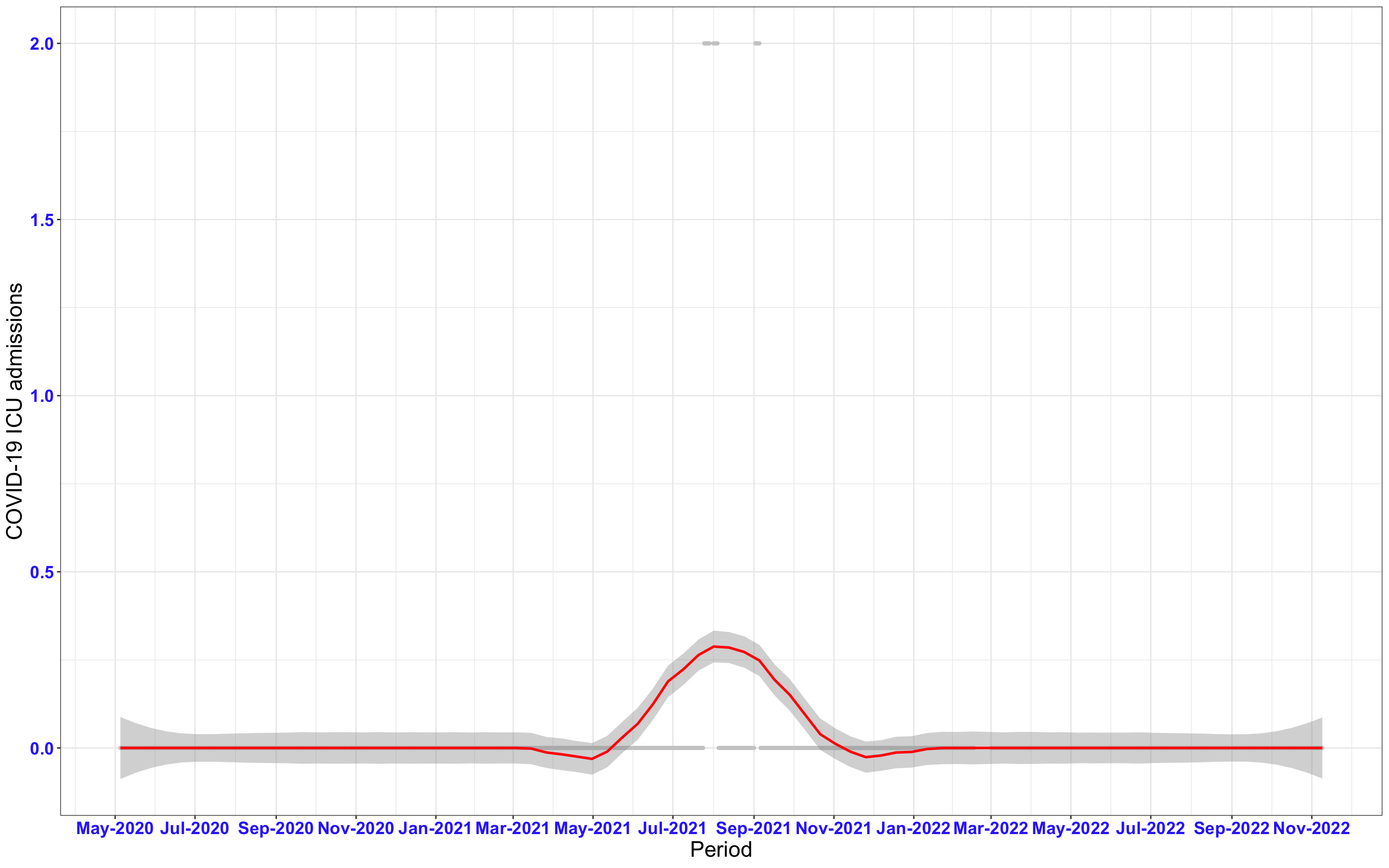
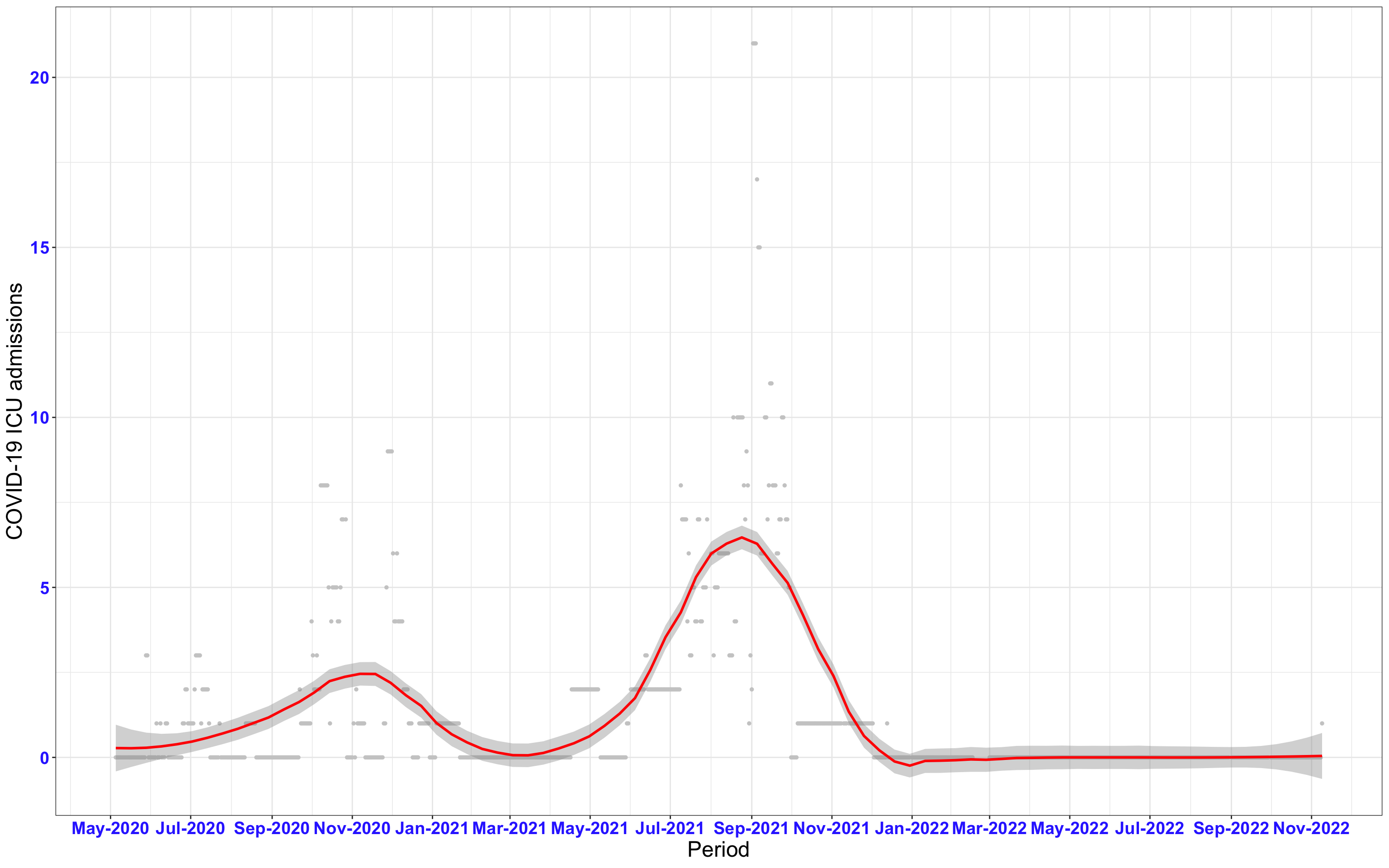
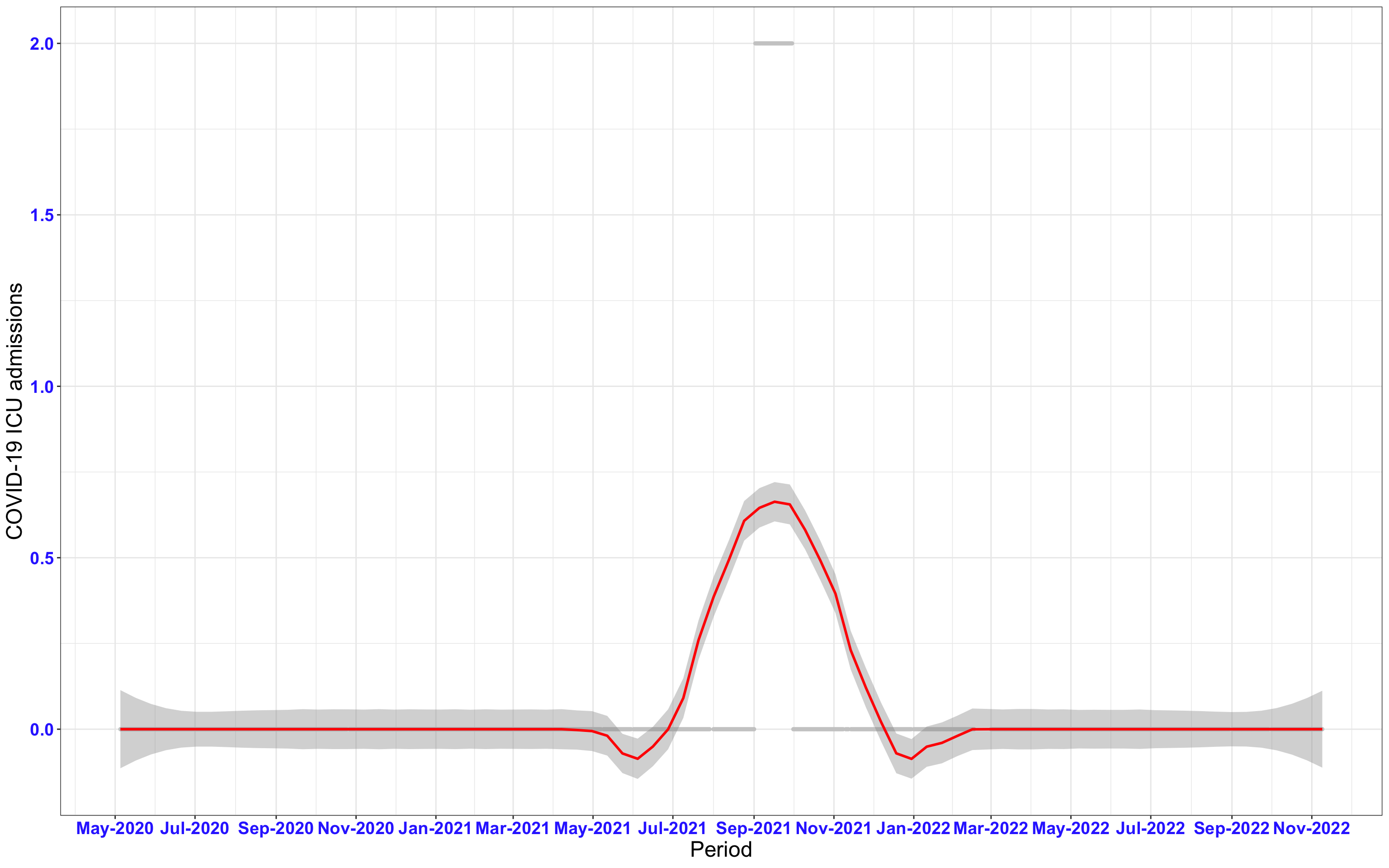
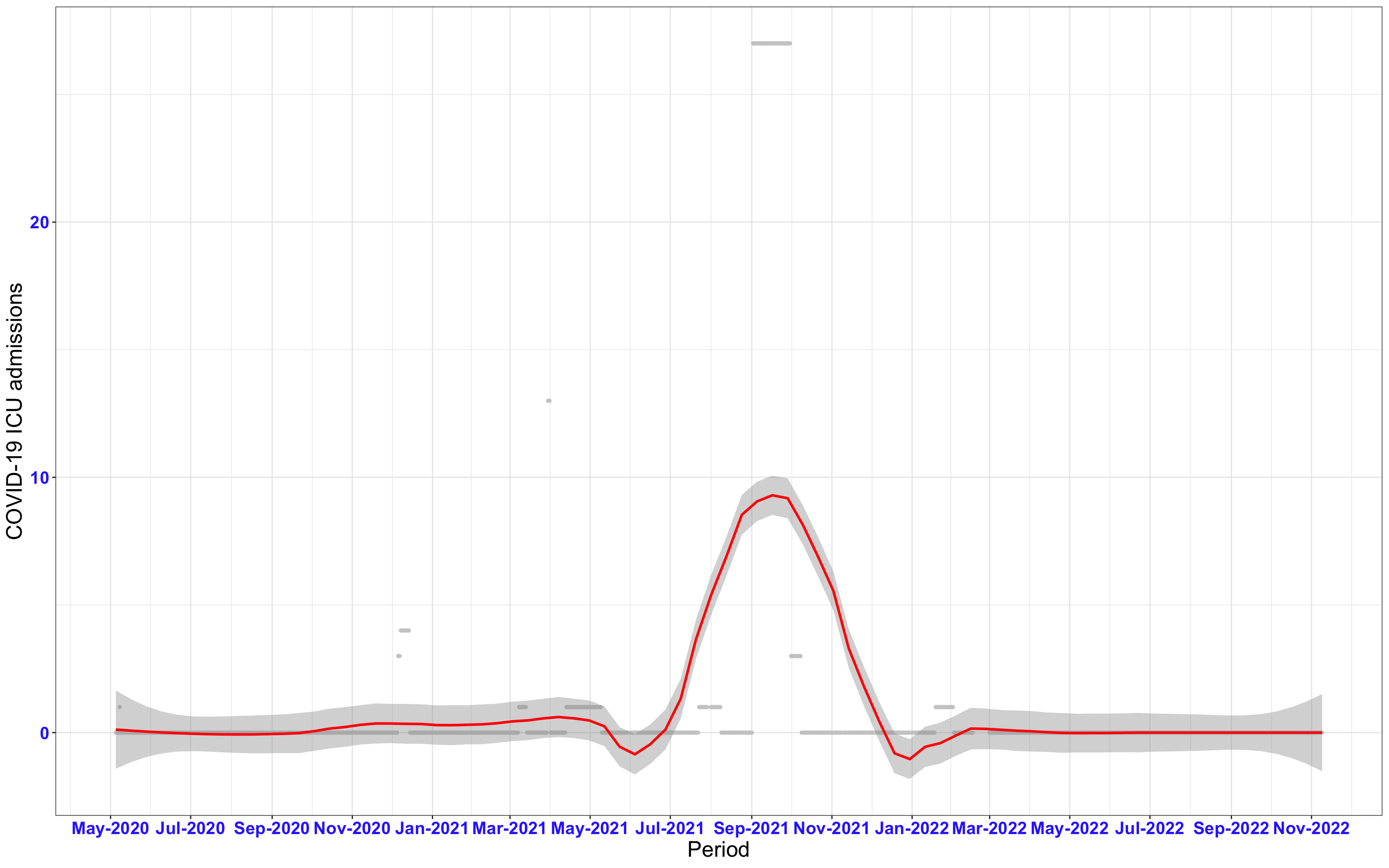
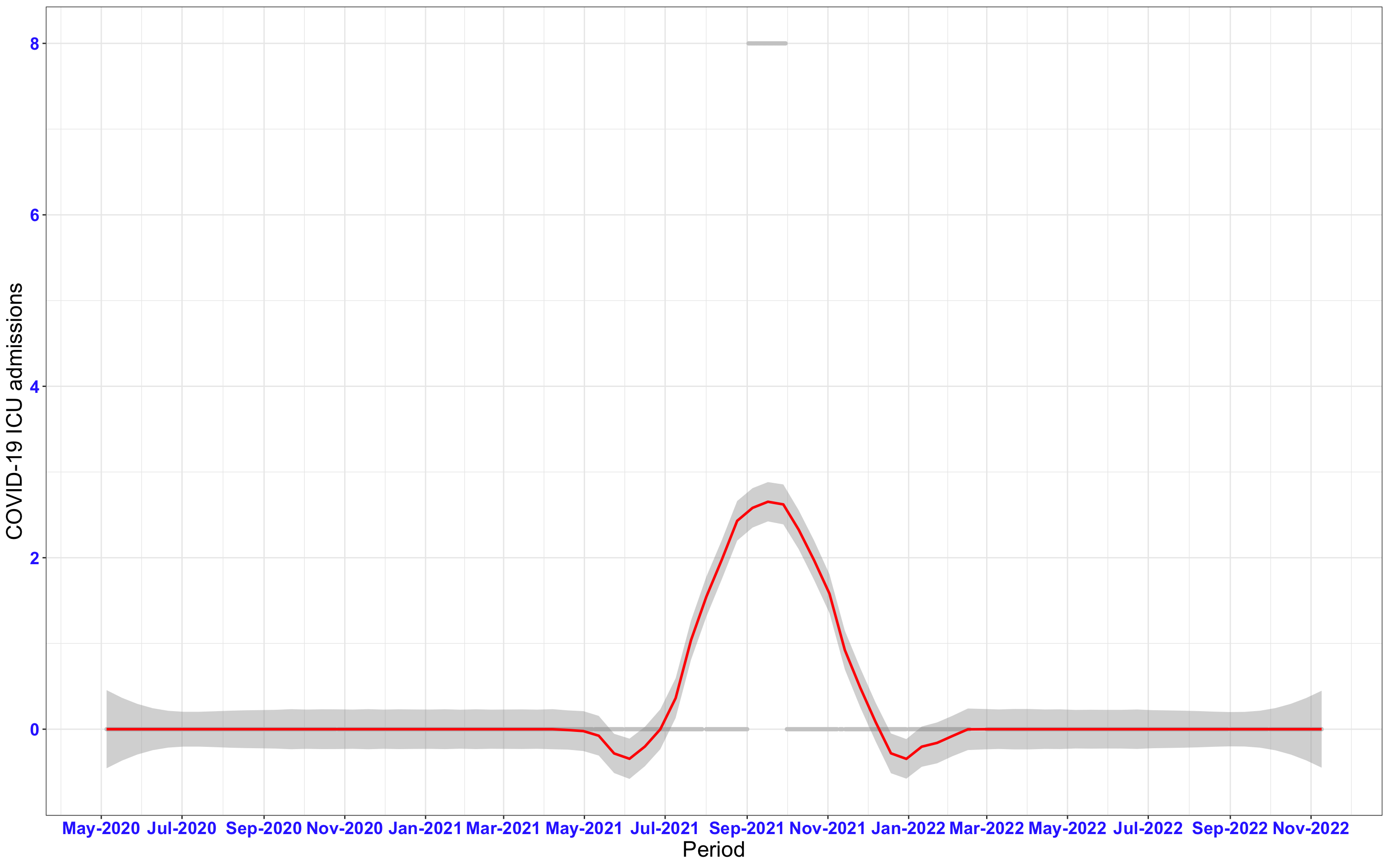
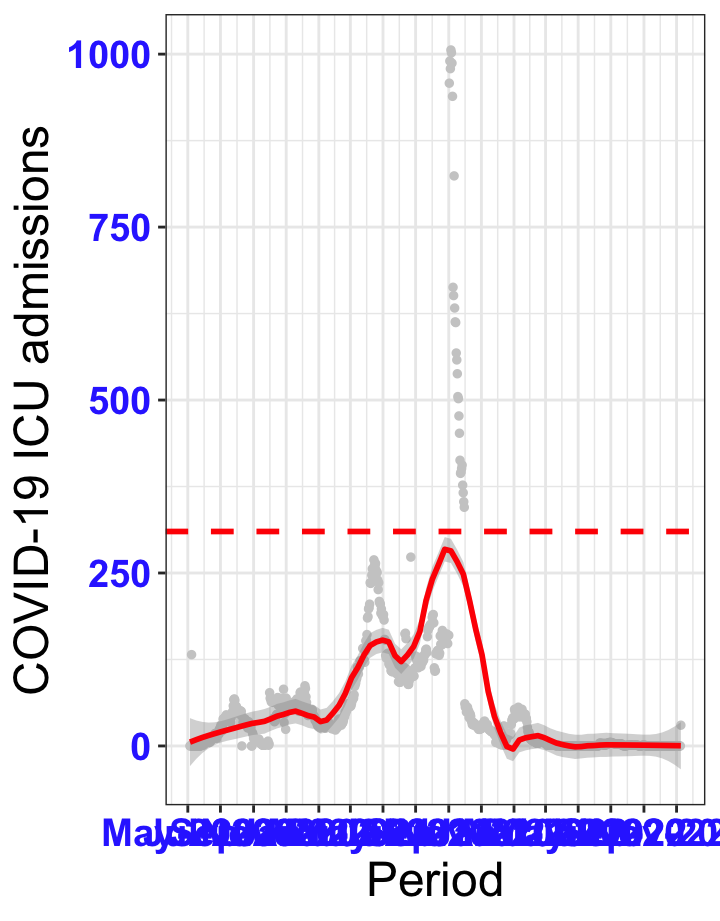
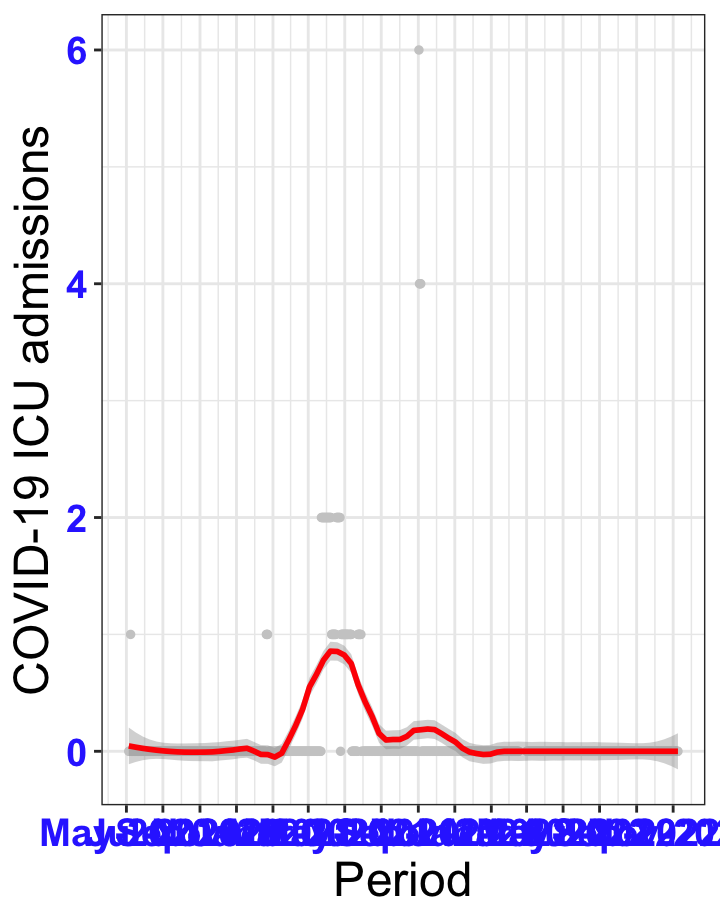
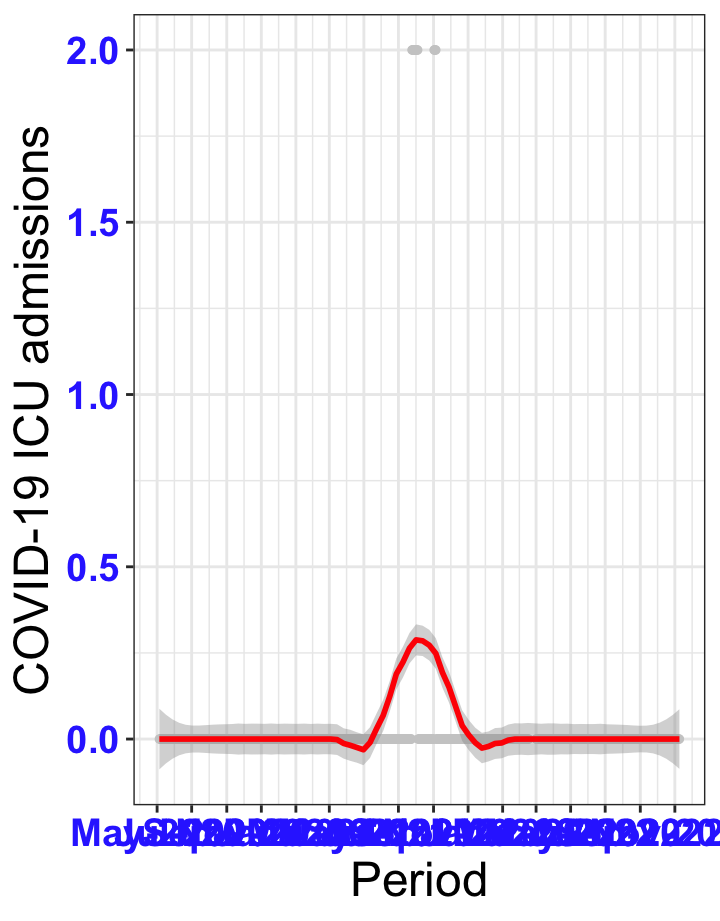
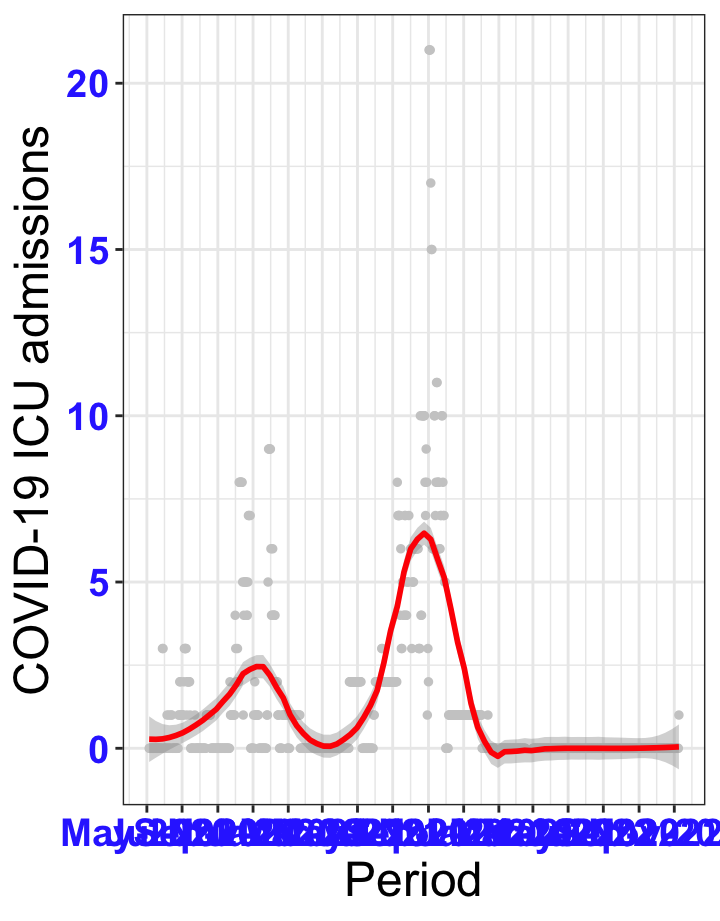
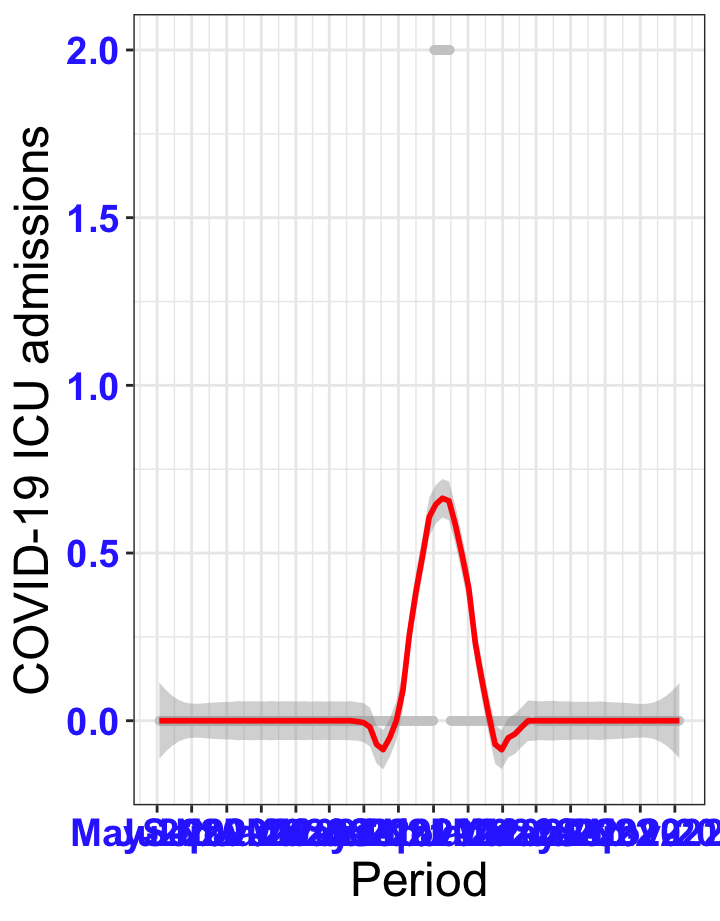
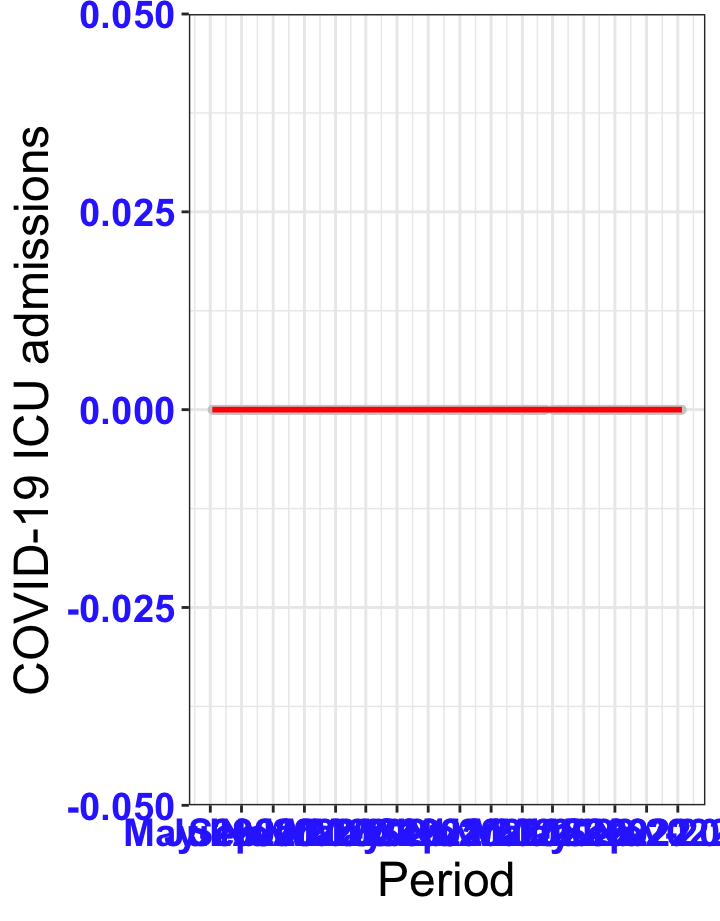
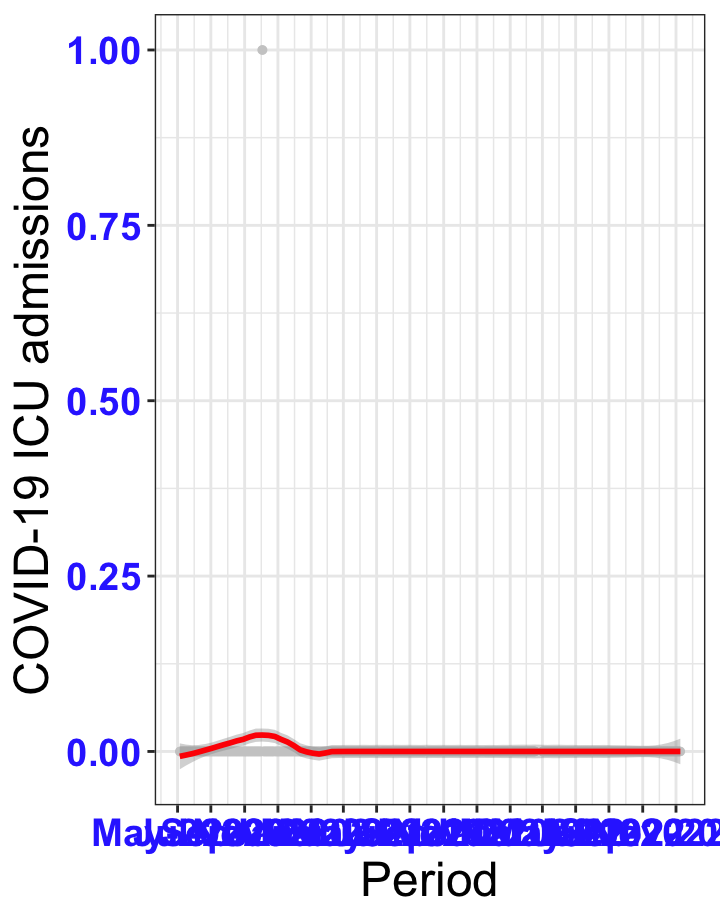
# ICU admissions {data-navmenu="ICU admissions"}
```{r,fig.width=16, fig.height=10}
p2<-ggplot(data1[data1$date>as.Date("2020-05-04"),], aes(x=date,y=icu))+geom_point(color="grey80", size=1)+theme_bw()+geom_smooth(span=0.3, colour="red", oob="squish")+scale_x_date(date_labels ="%b-%Y" ,date_breaks = "2 months")+labs(x="Period", y="COVID-19 ICU admissions")+theme(text=element_text(size=18 ), axis.text = element_text(color = "#3339FF", face = "bold"))+geom_hline(yintercept = 310, linetype=2, color="red", size=1)
p2
```
# ICU admissions {data-navmenu="Visualisation"}
```{r,,fig.width=16, fig.height=10, message=F}
plot2<-ggplot(county_admissions, aes(fill=icu, label=Name))+geom_sf()+theme_void()+scale_fill_gradient(high="#f03b20", low="#ffeda0", na.value = "grey80")+labs(fill="",title="Average number of ICU admissions over the last one week")
ggplotly(plot2)
```
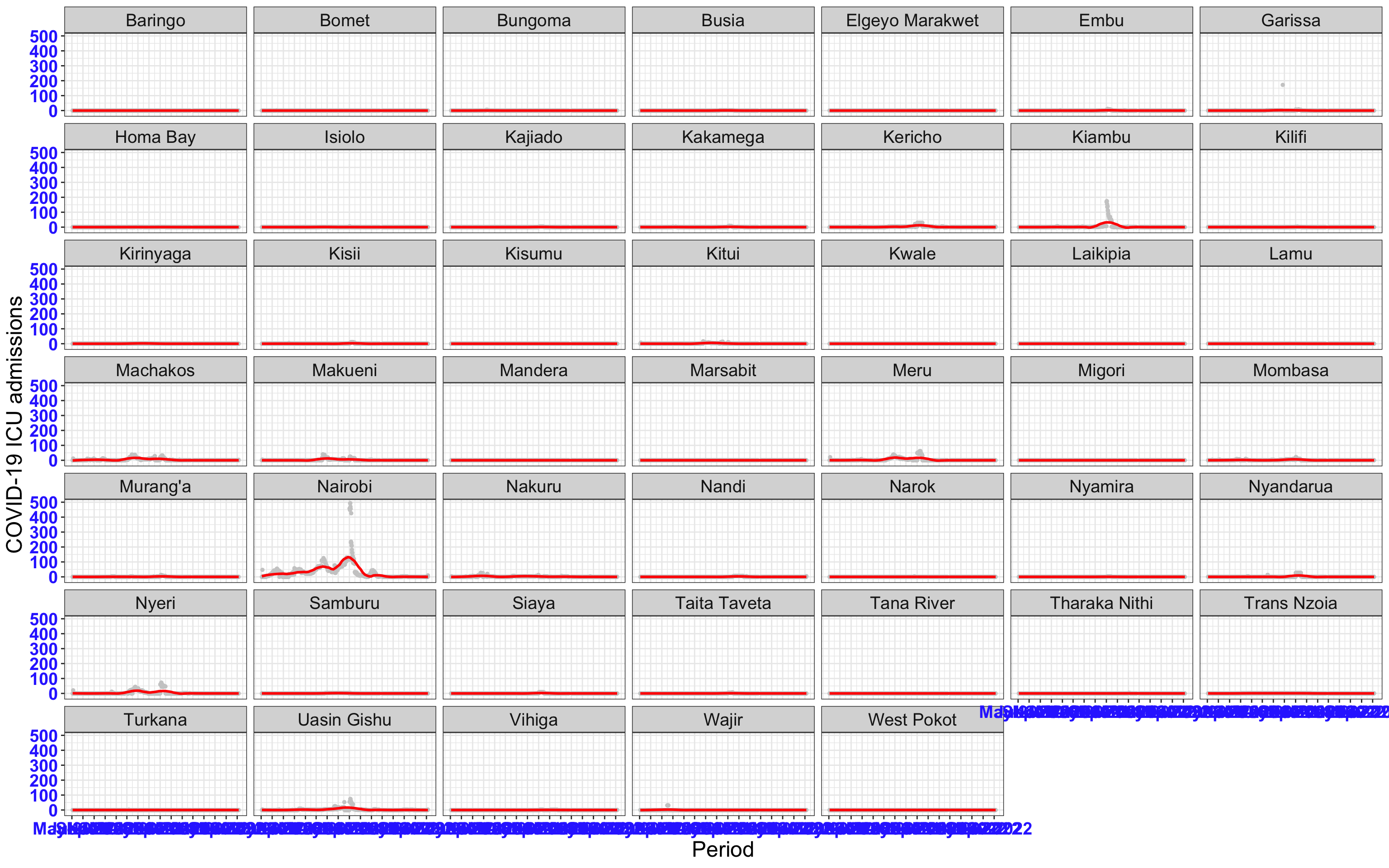
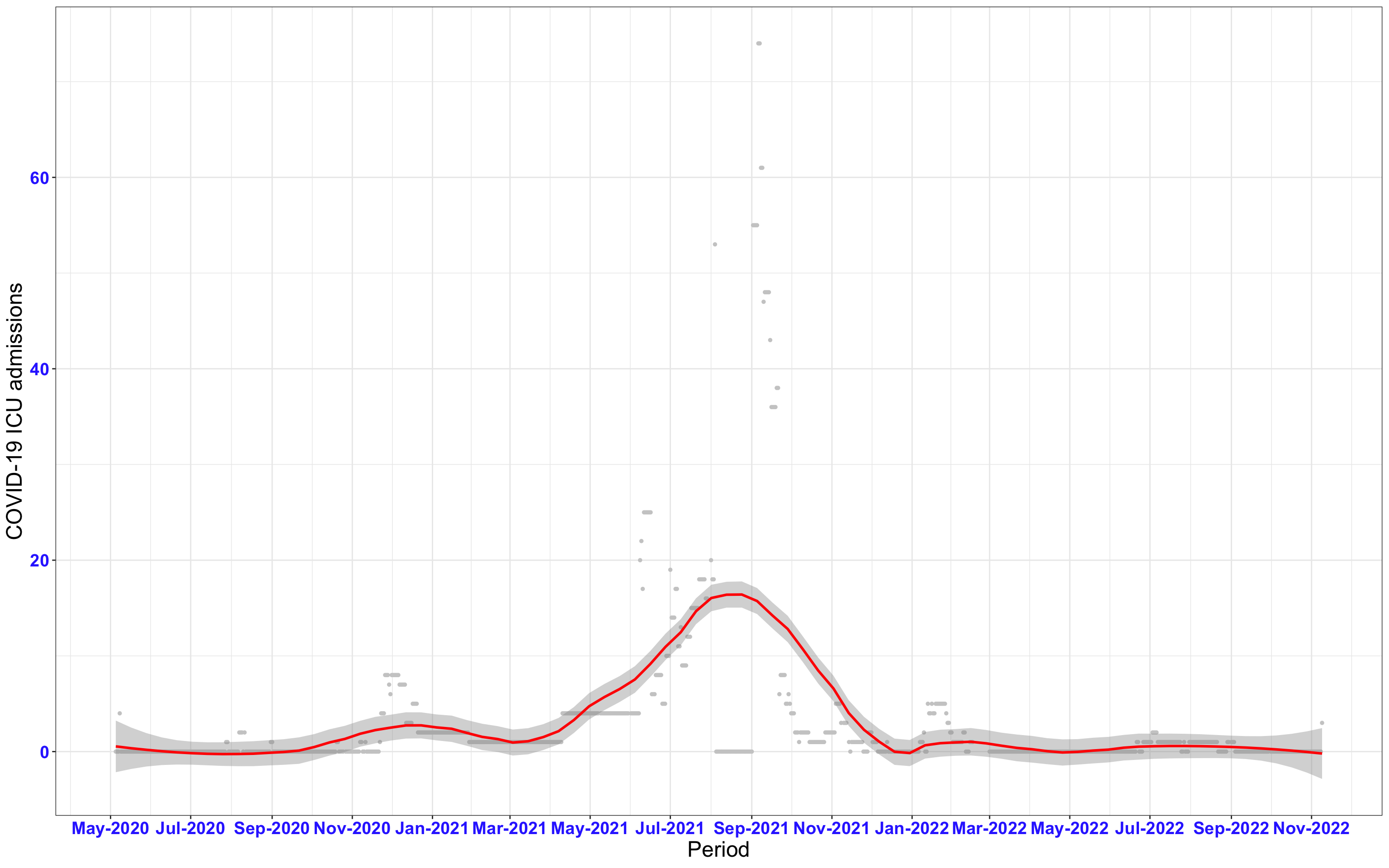
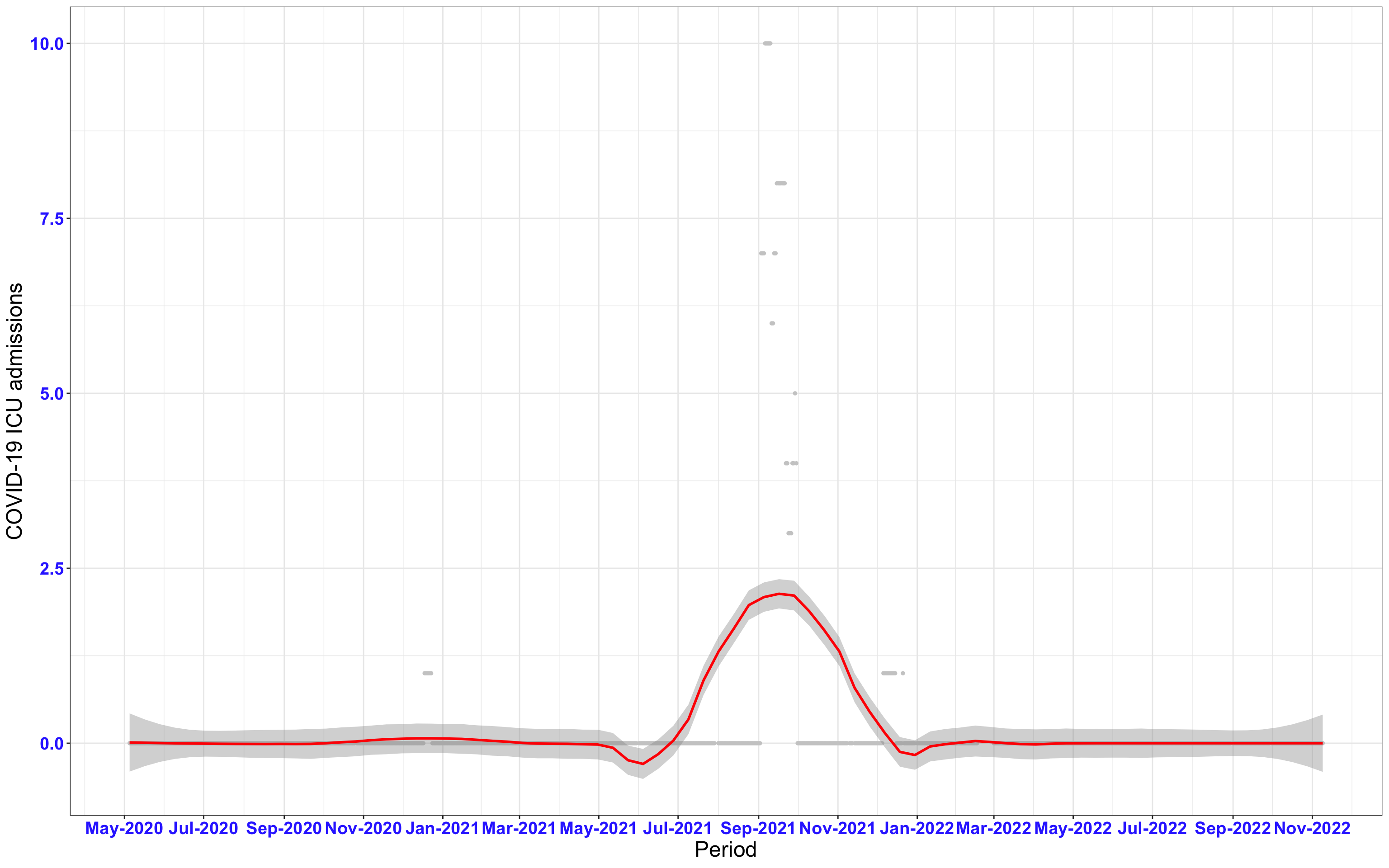
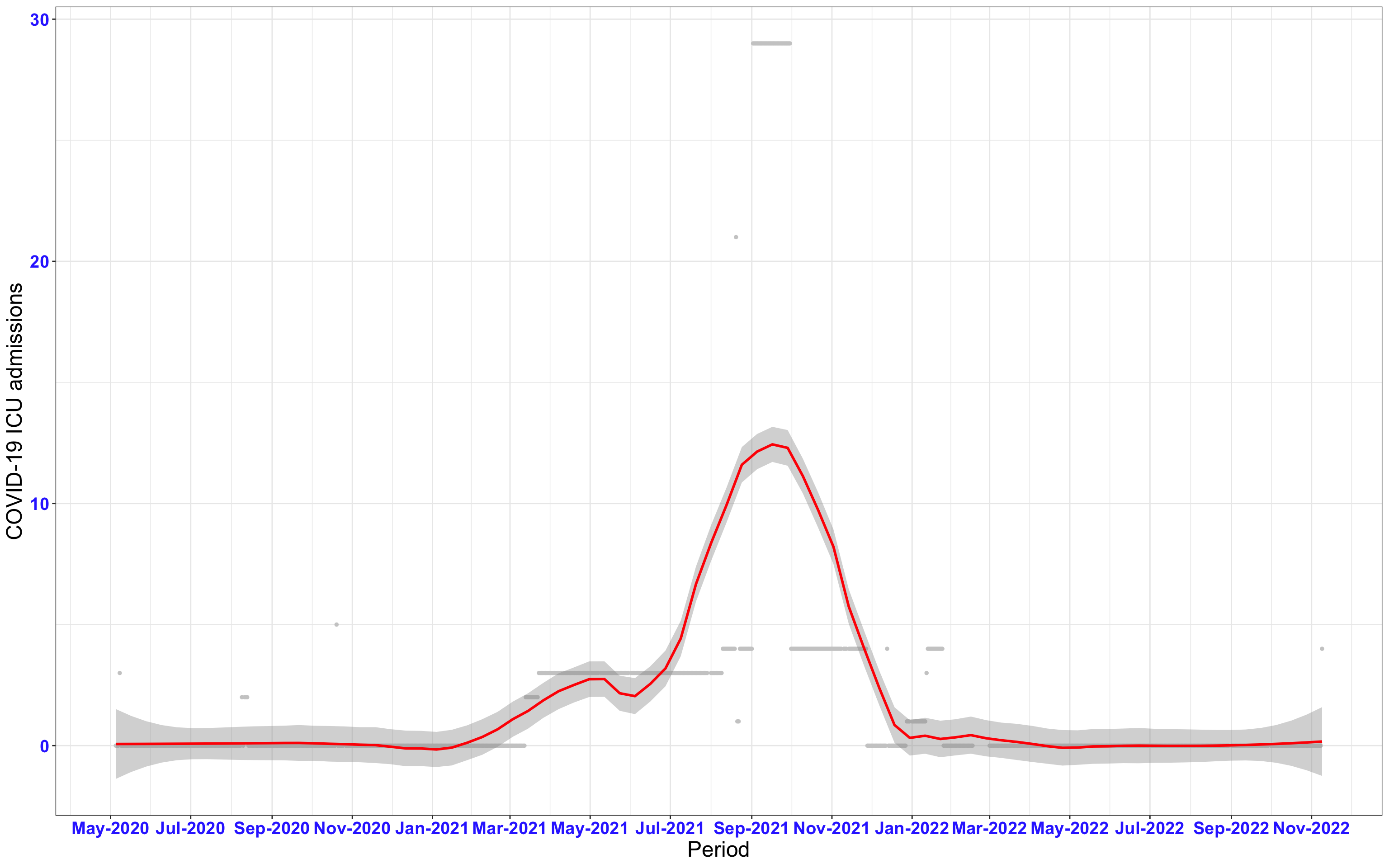
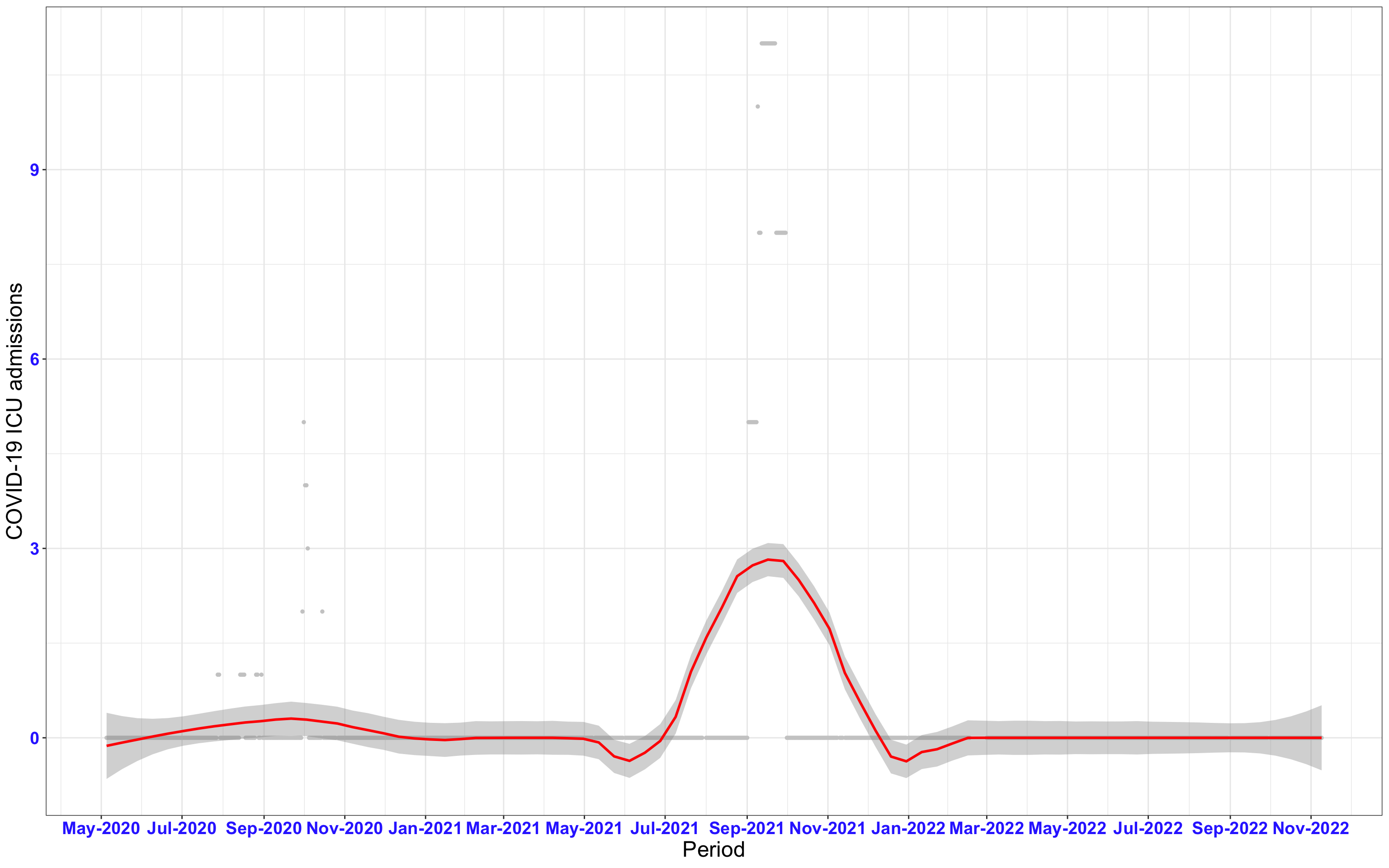
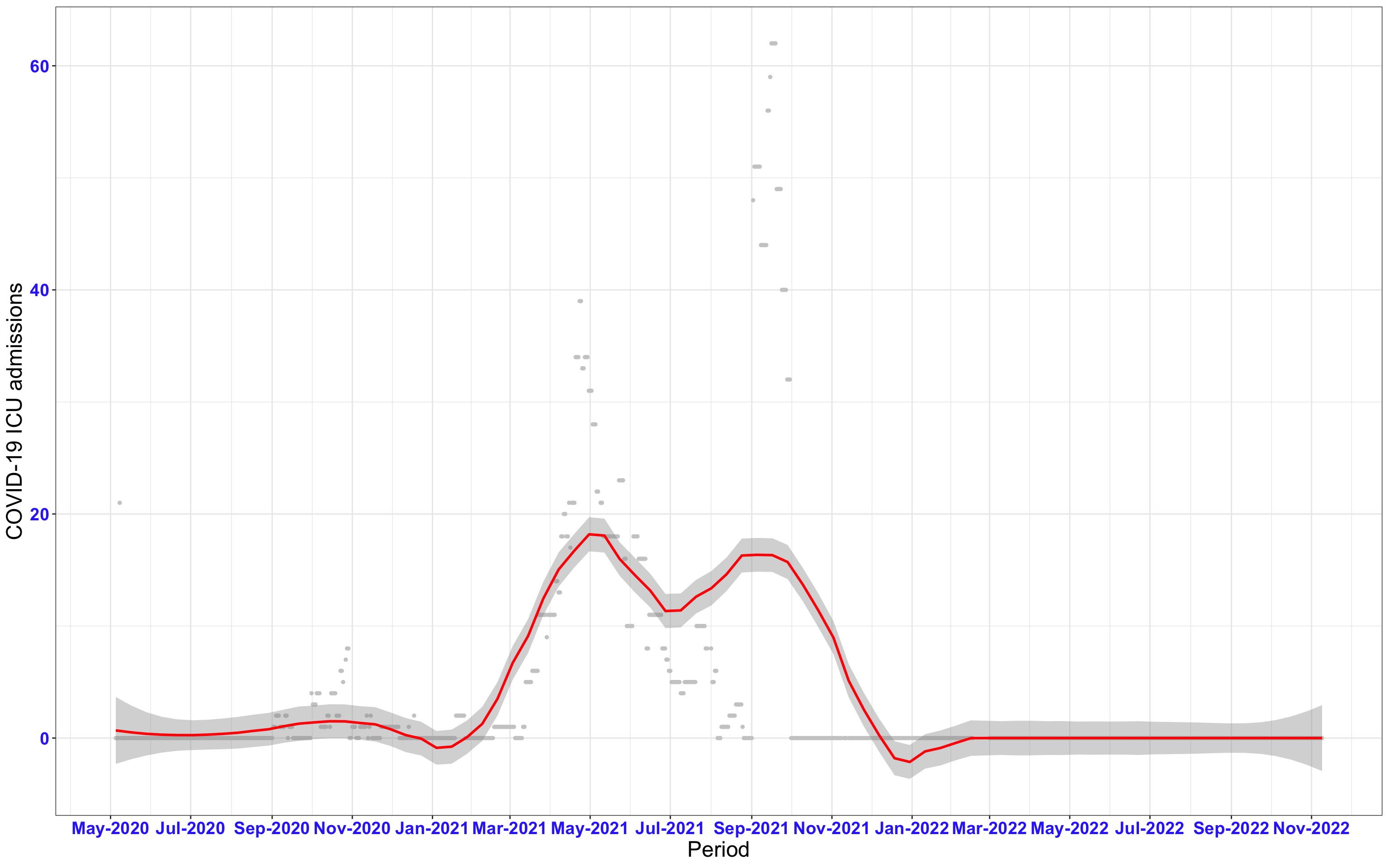
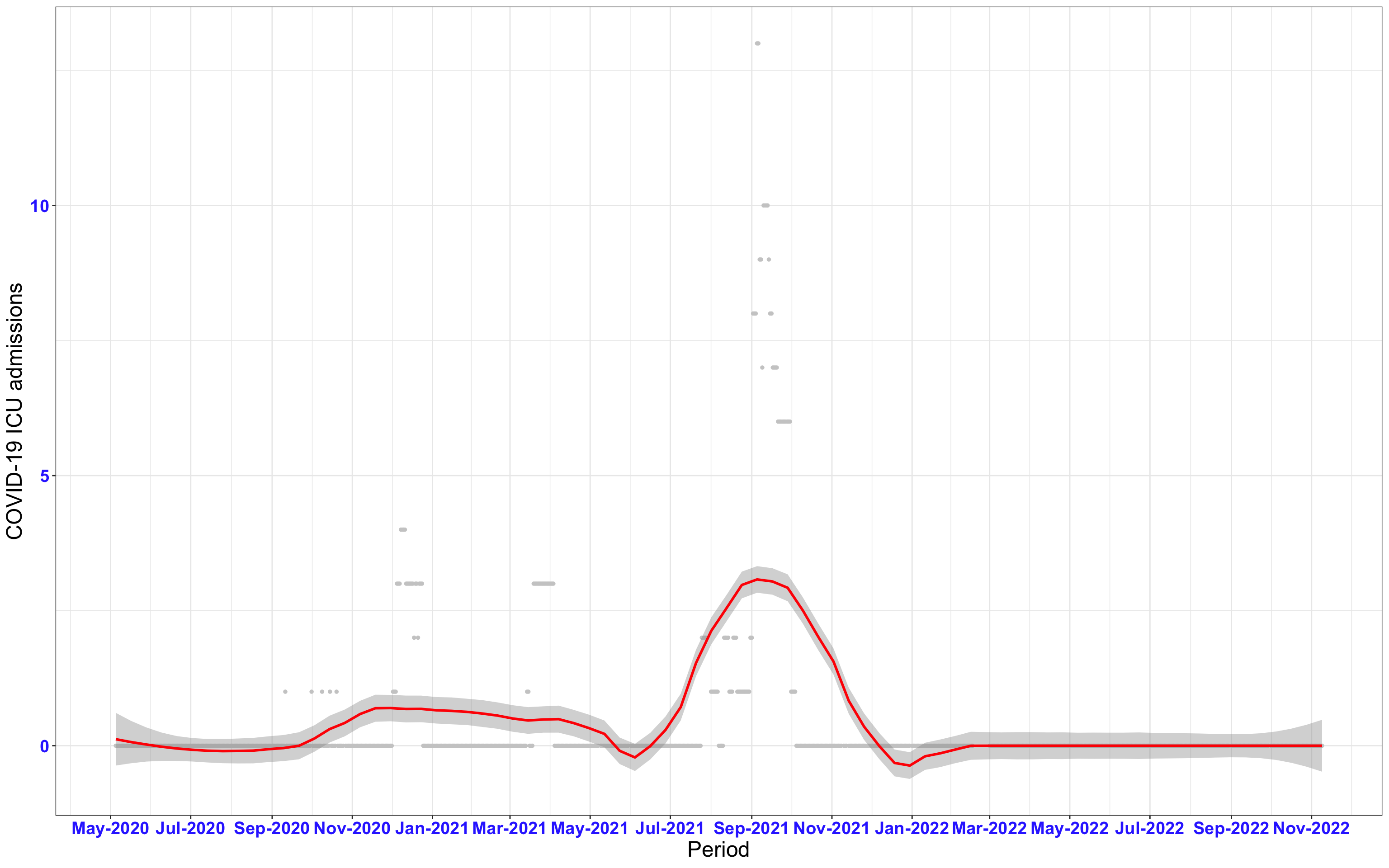
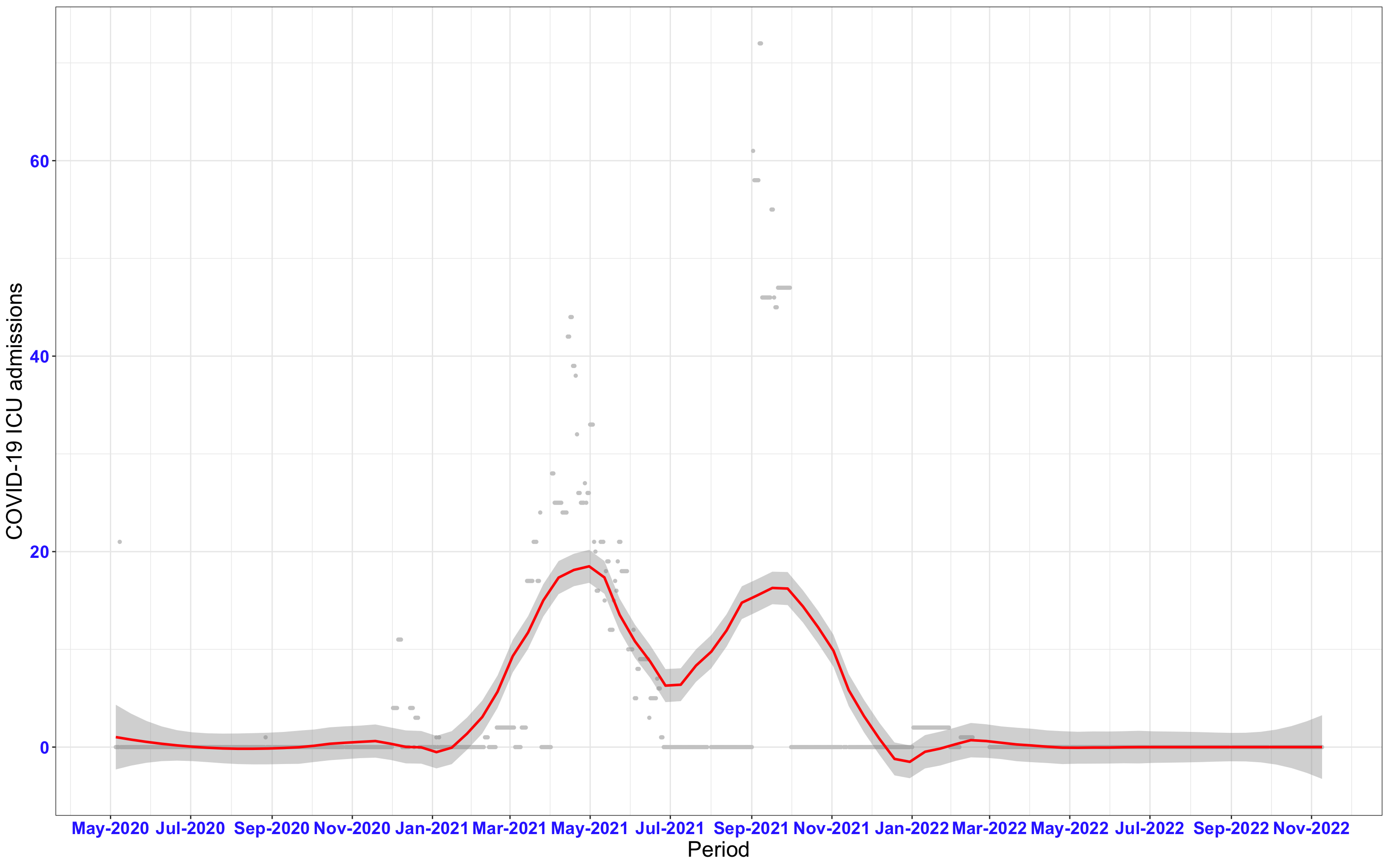
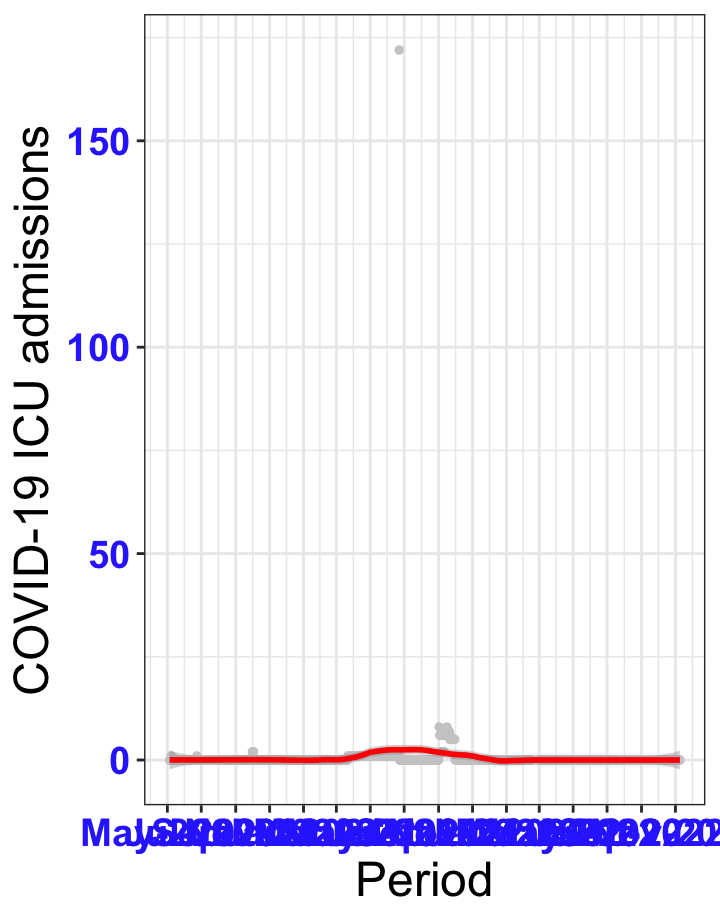
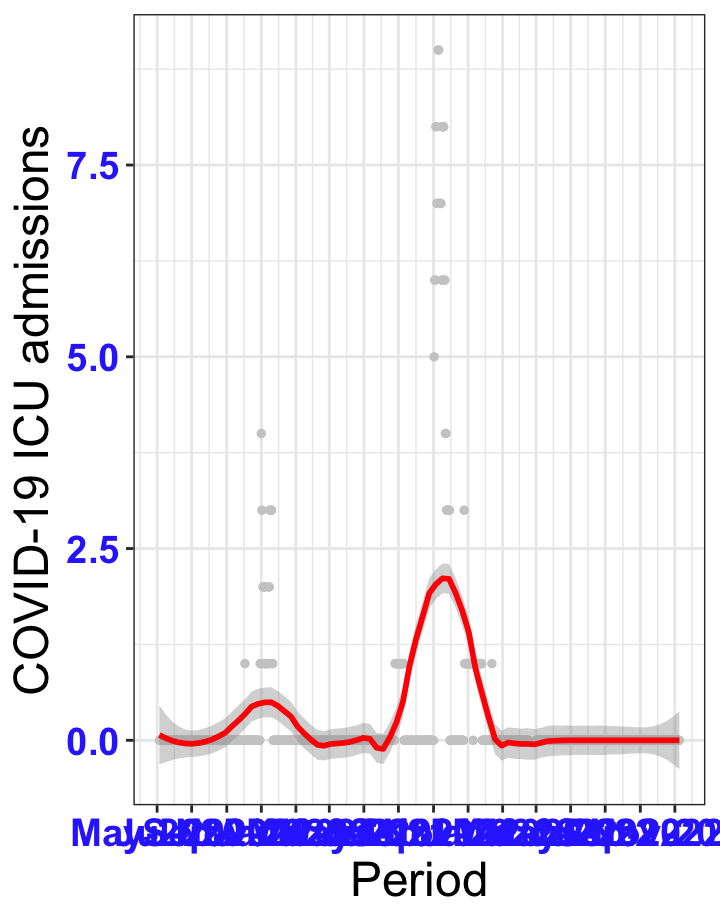
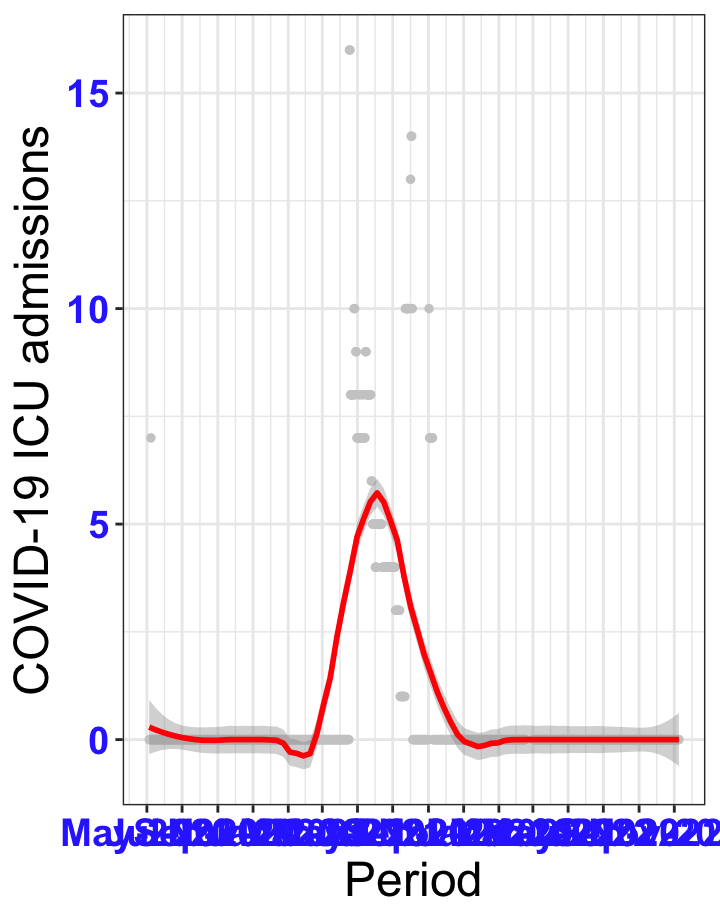
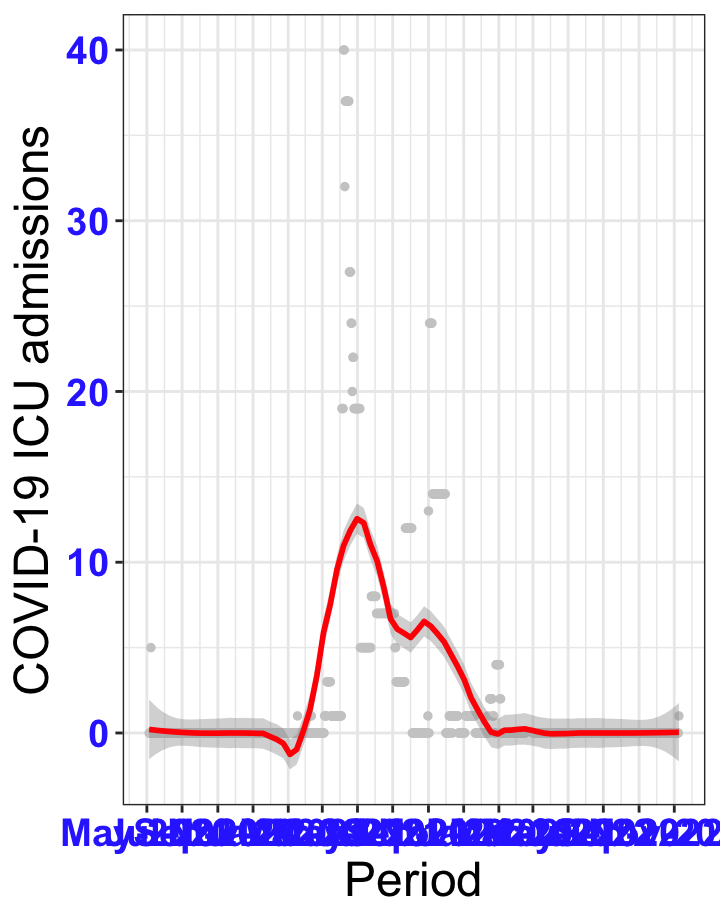
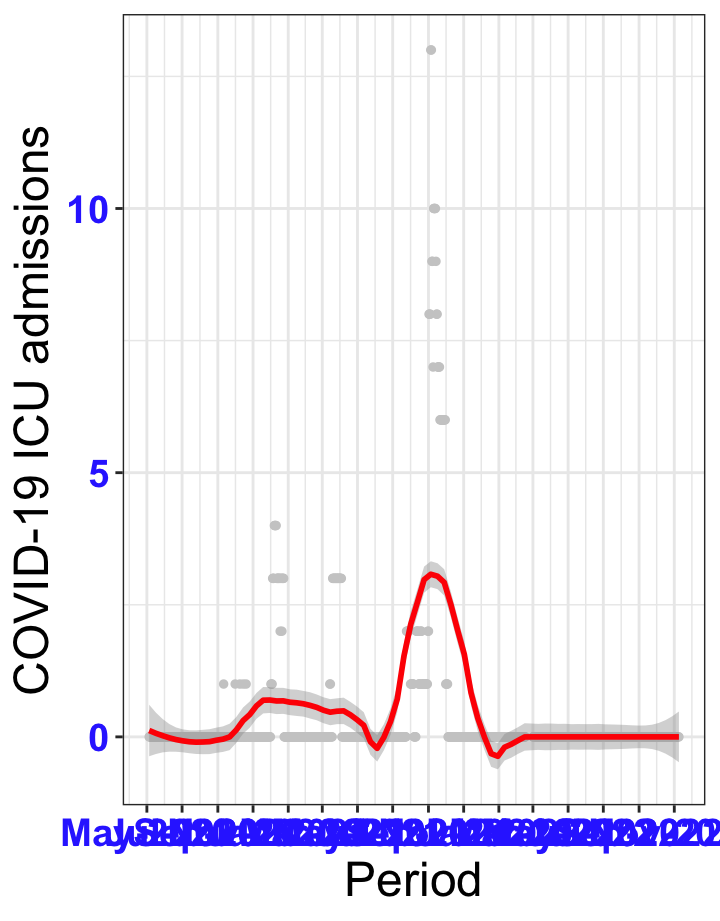
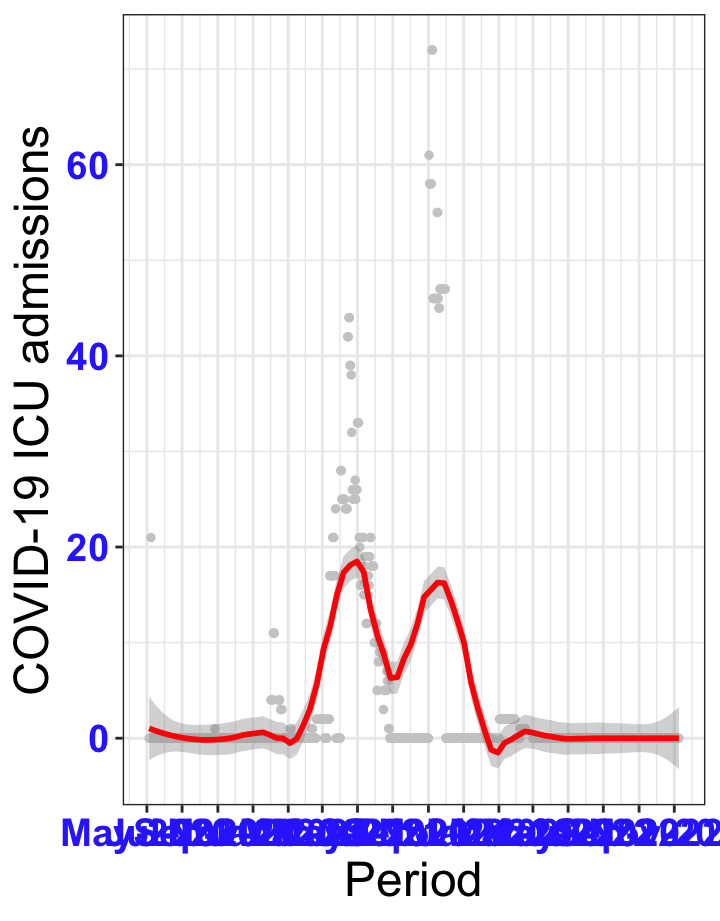
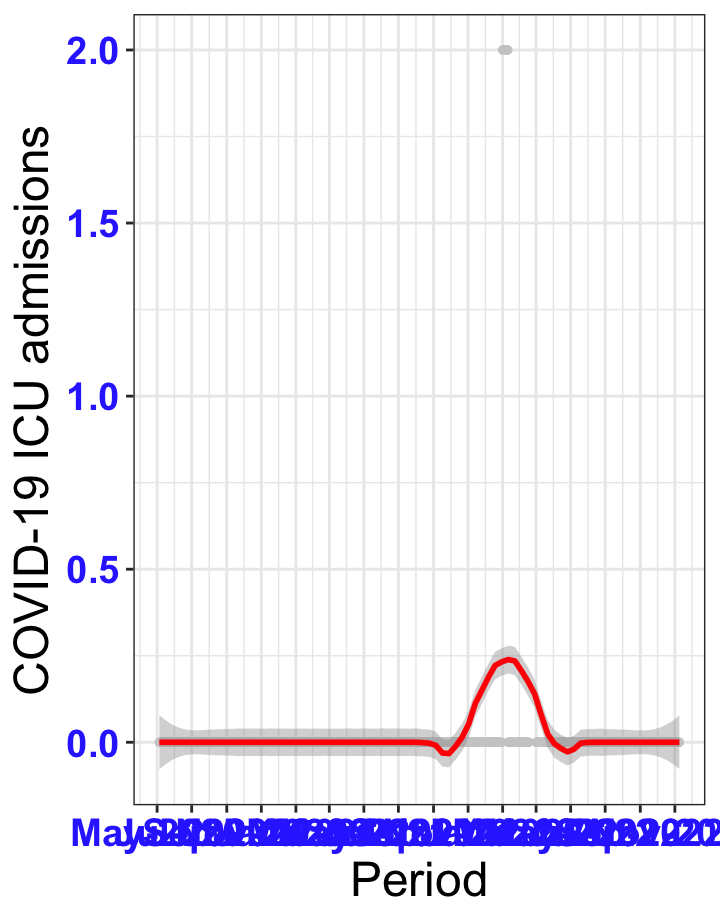
# All counties {data-navmenu="ICU admissions"}
```{r, fig.width=16, fig.height=10}
ggplot(data_county[data_county$date>as.Date("2020-05-04"),],aes(x=date,y=icu))+
geom_point(color="grey80", size=1)+facet_wrap(~county)+theme_bw()+geom_smooth(span=0.3, colour="red", oob="squish")+scale_x_date(date_labels ="%b-%Y" ,date_breaks = "2 months")+labs(x="Period", y="COVID-19 ICU admissions")+theme(text=element_text(size=18 ), axis.text = element_text(color = "#3339FF", face = "bold"))
```
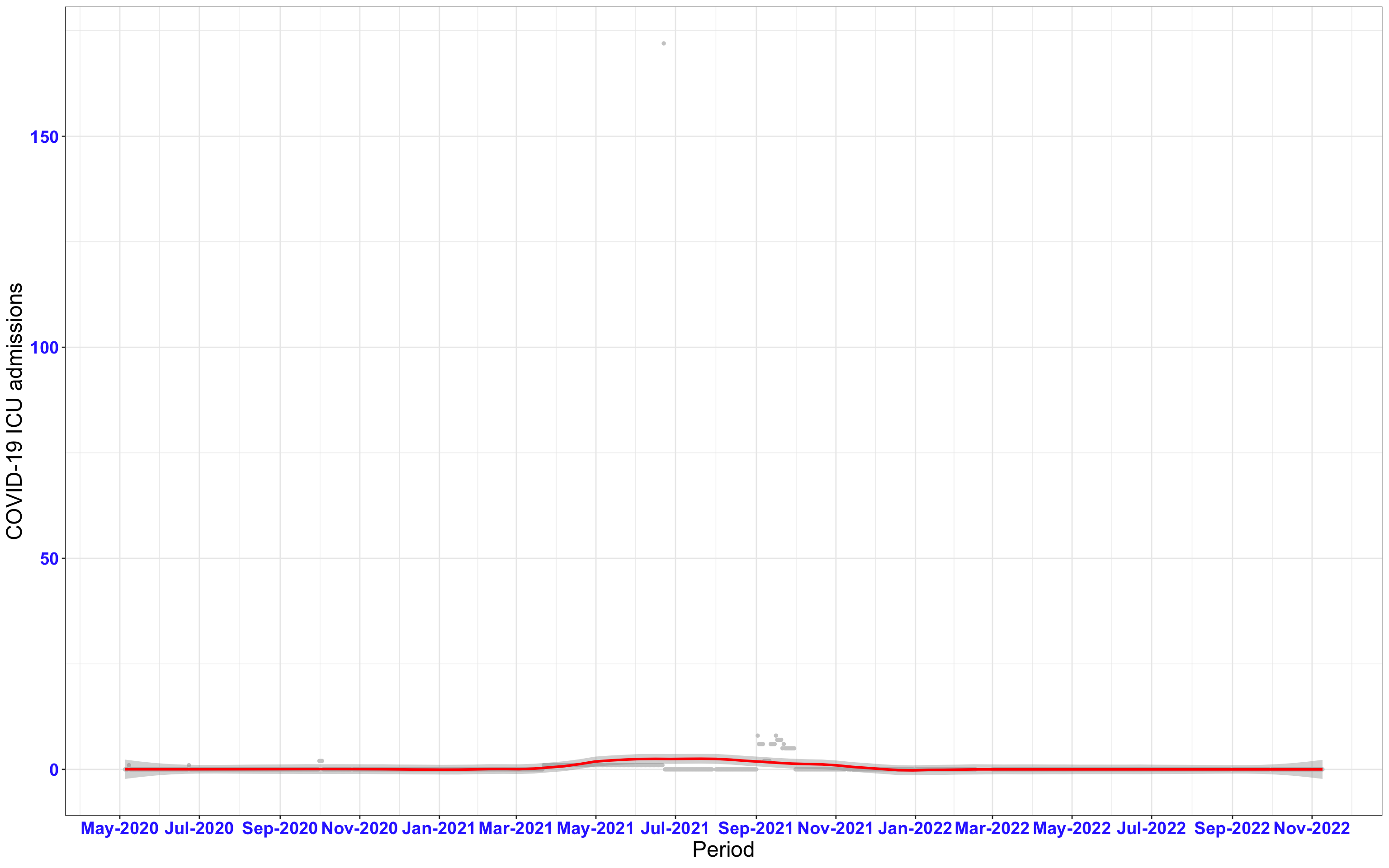
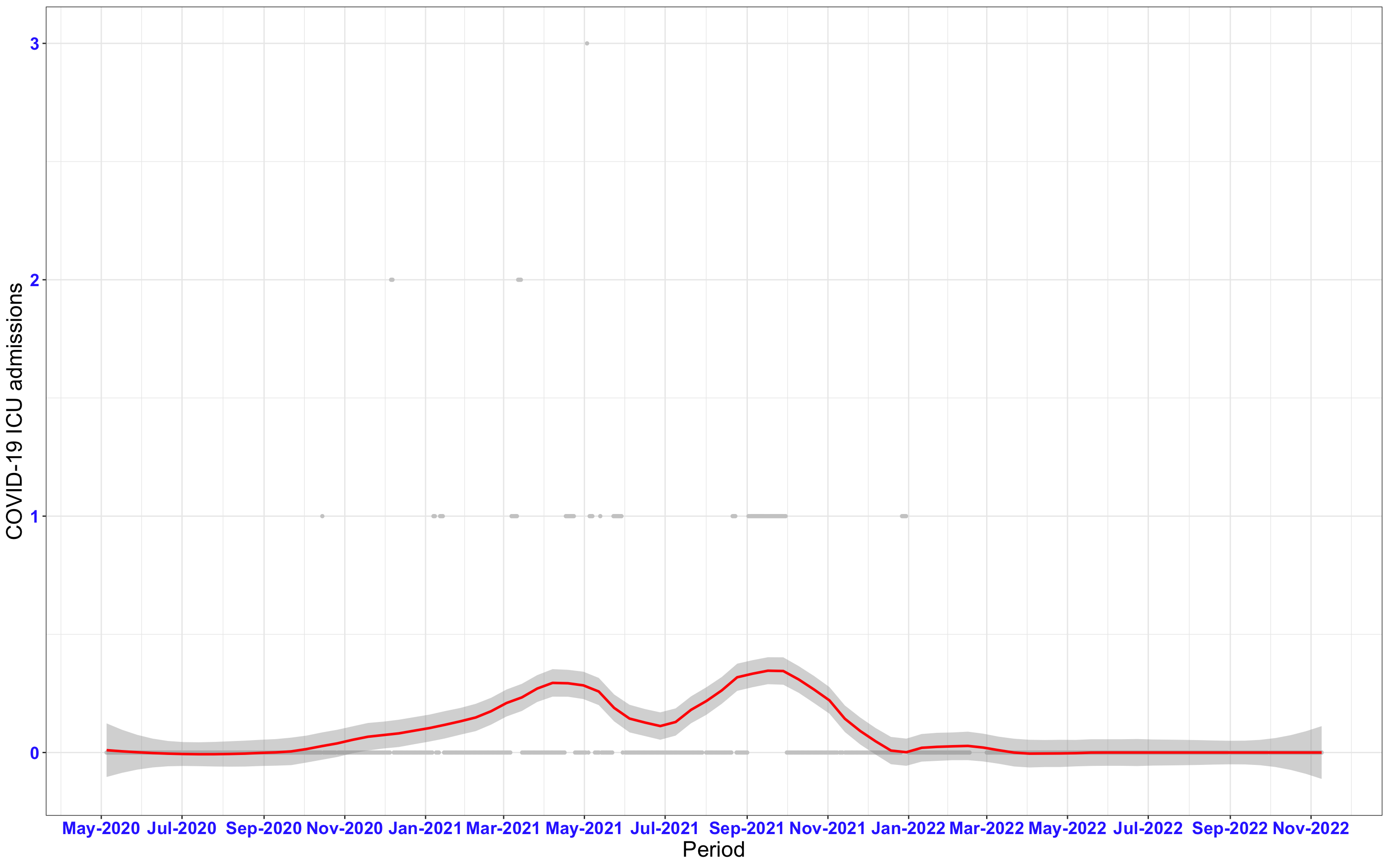
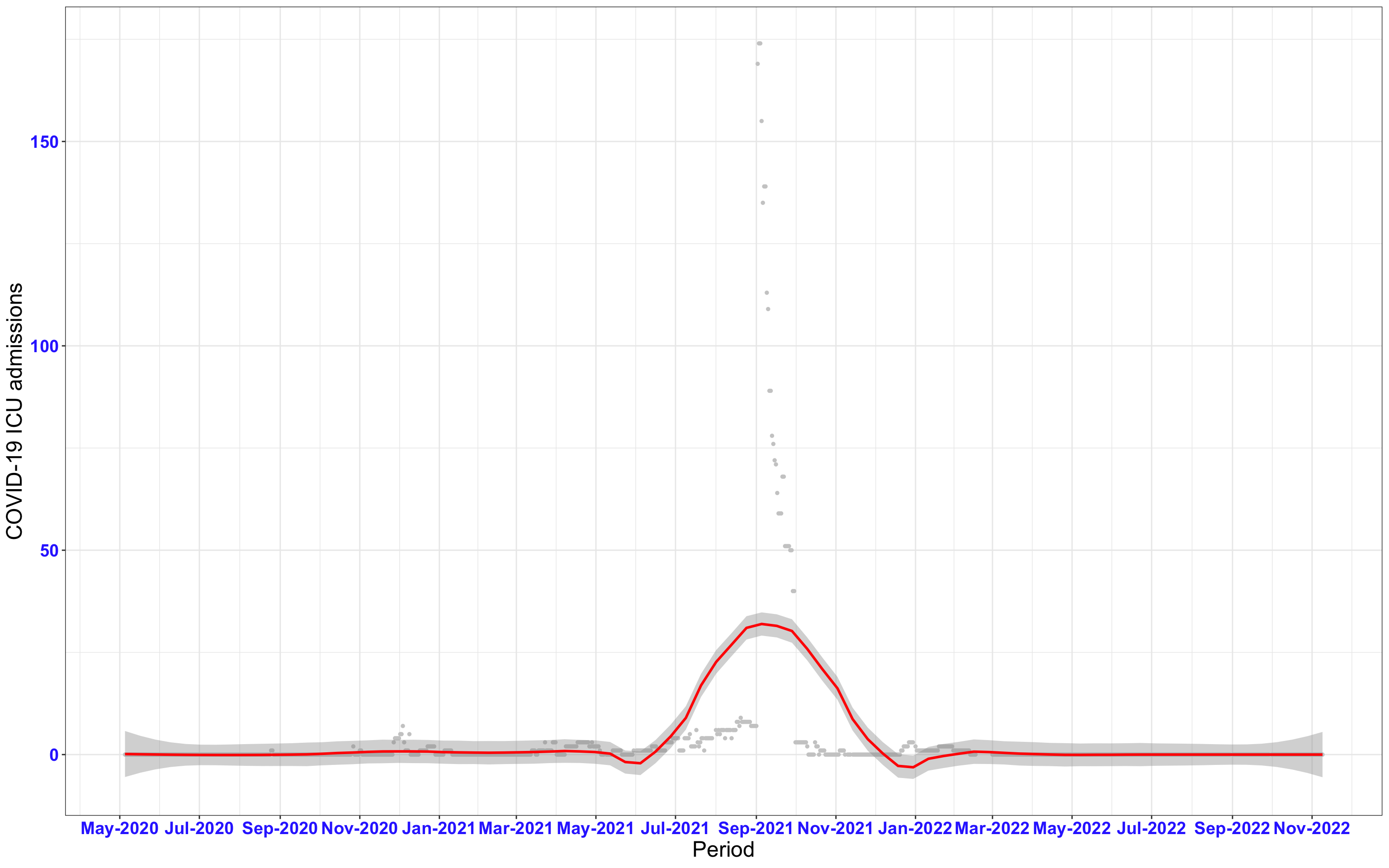
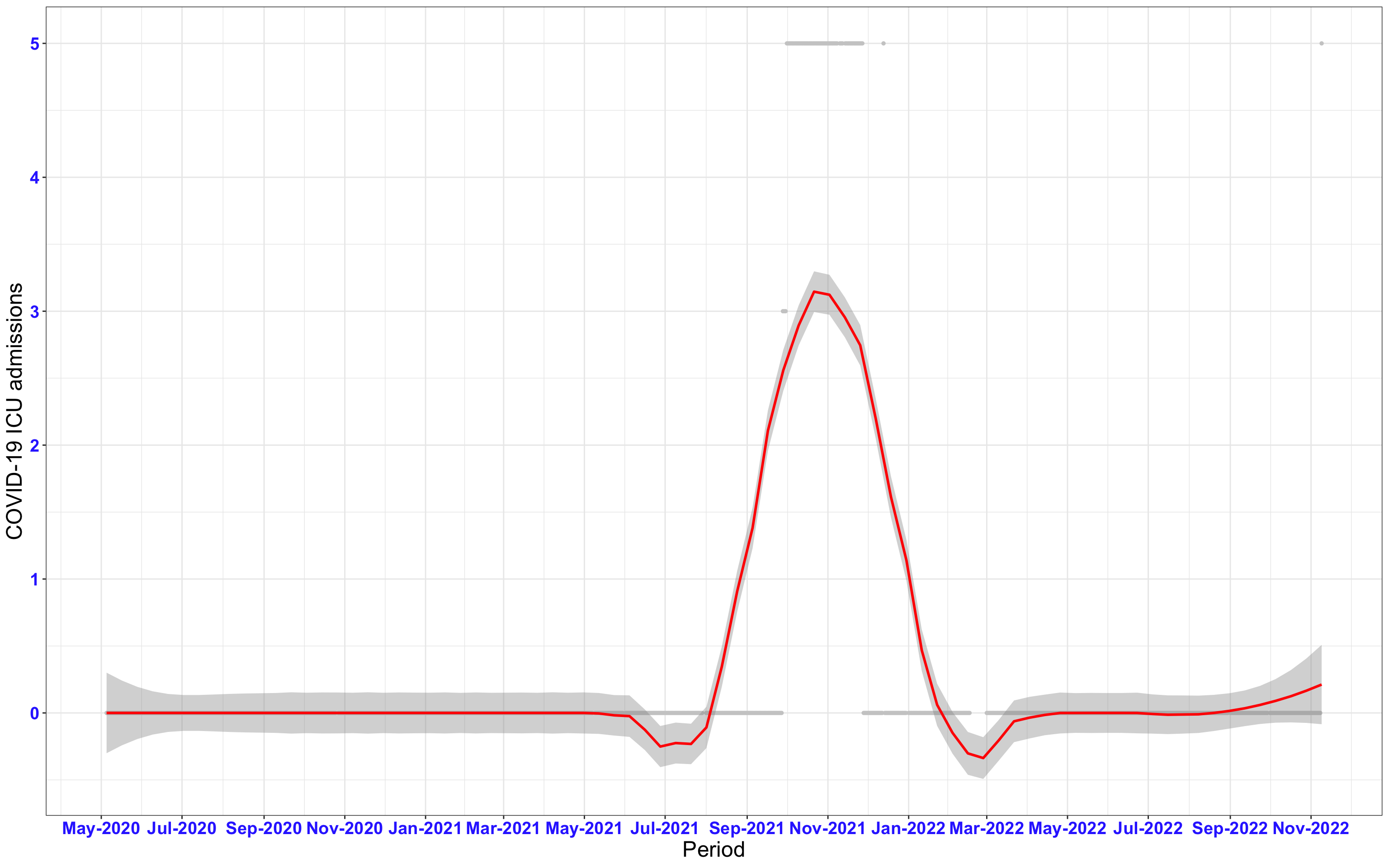
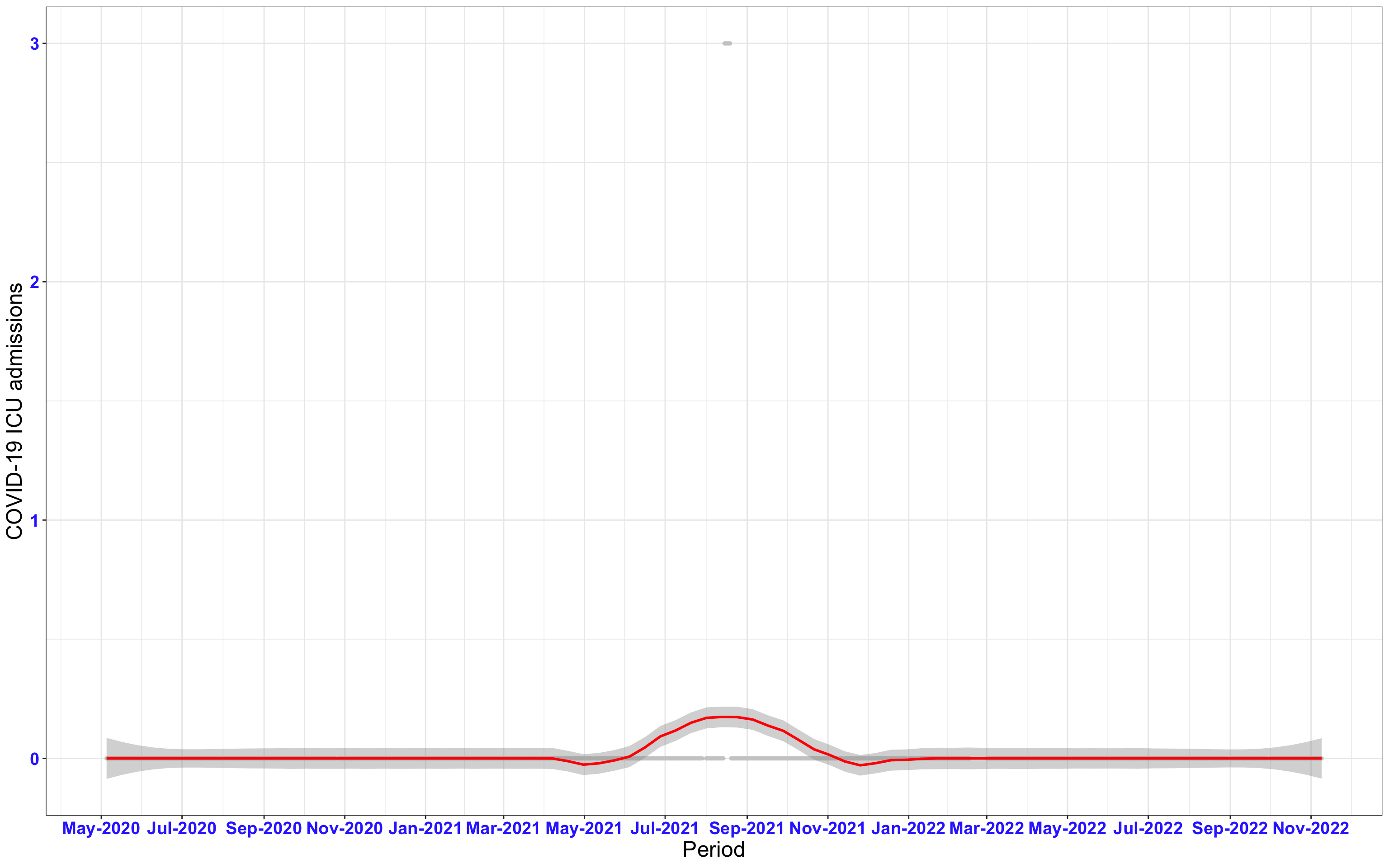
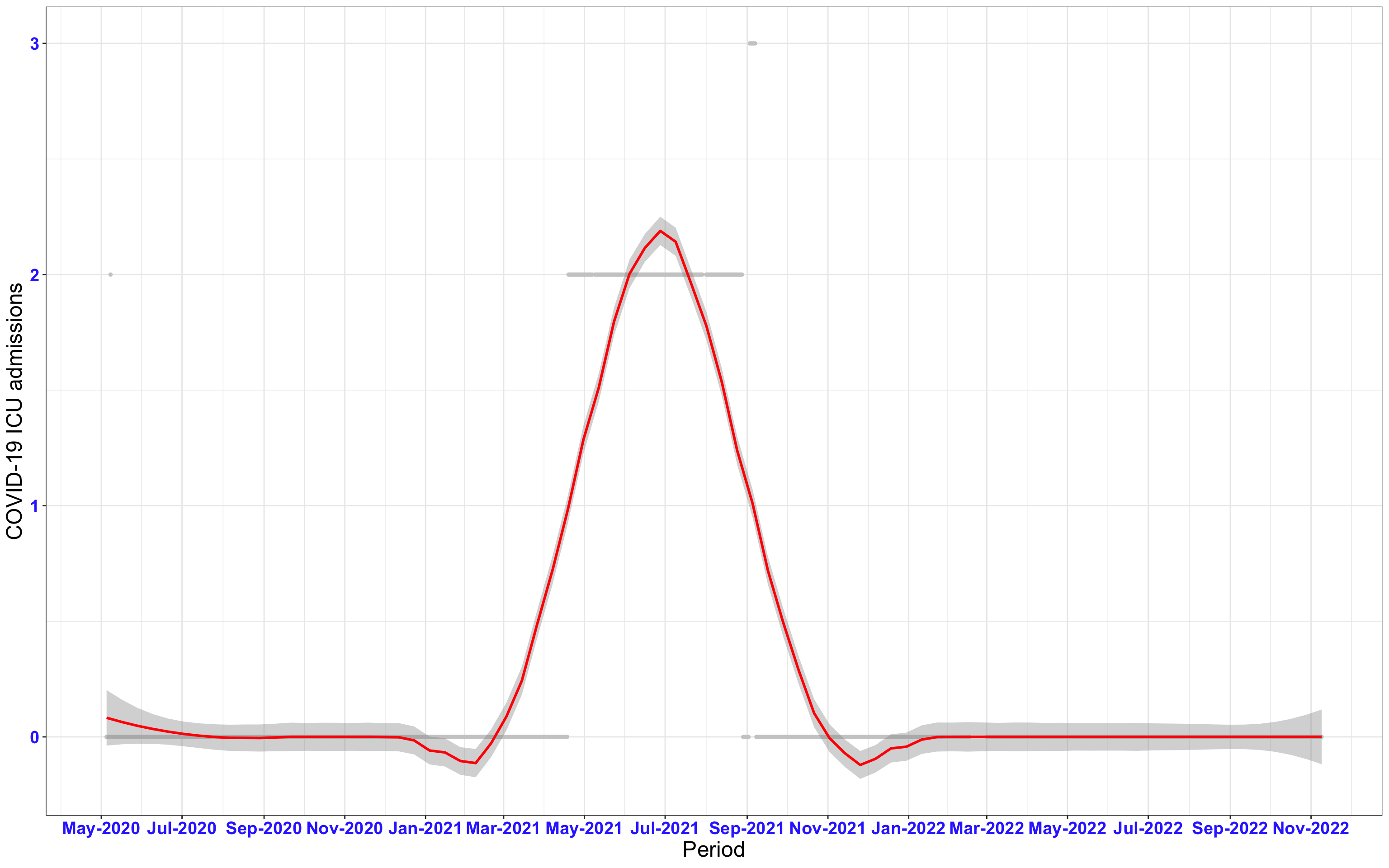
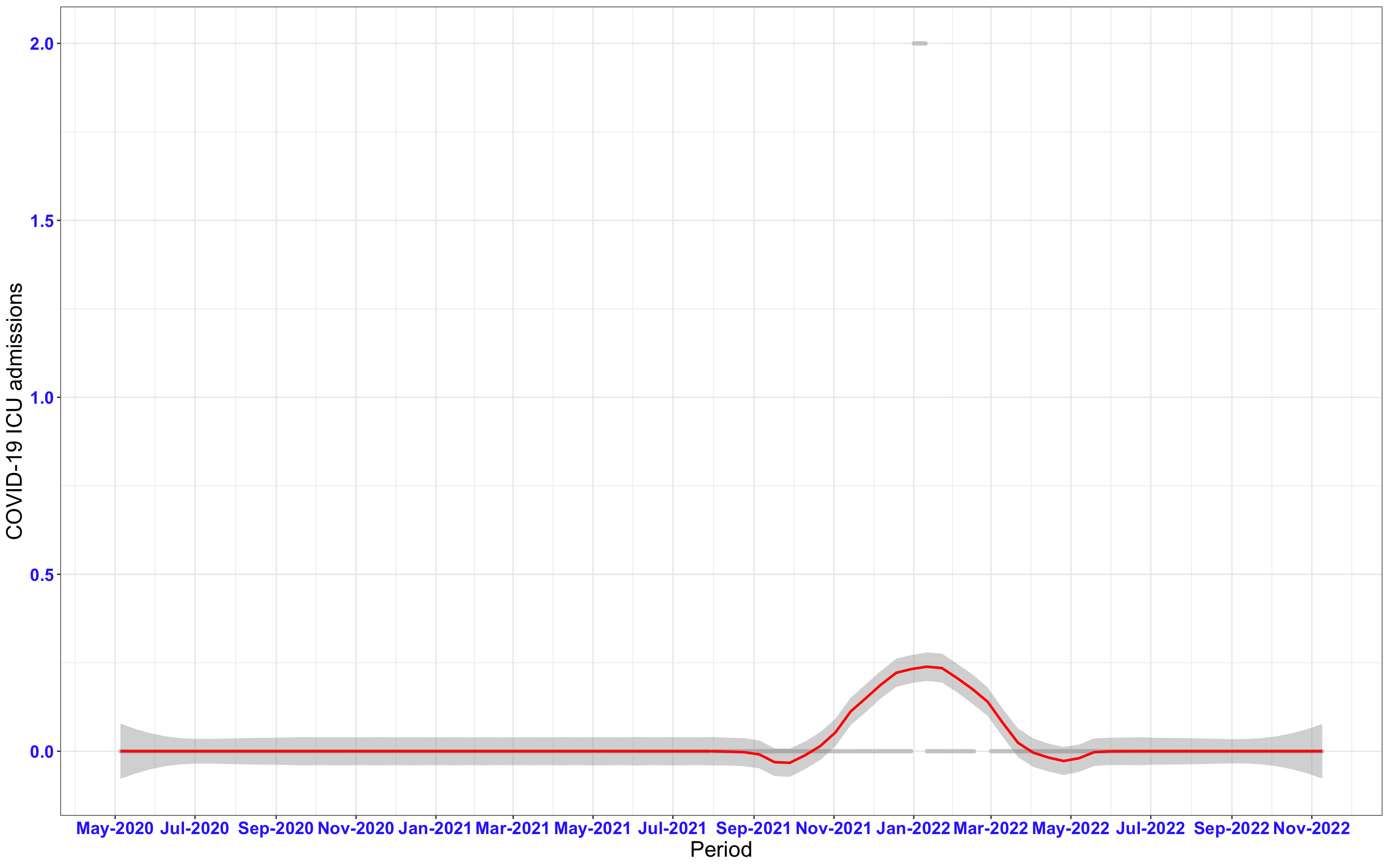
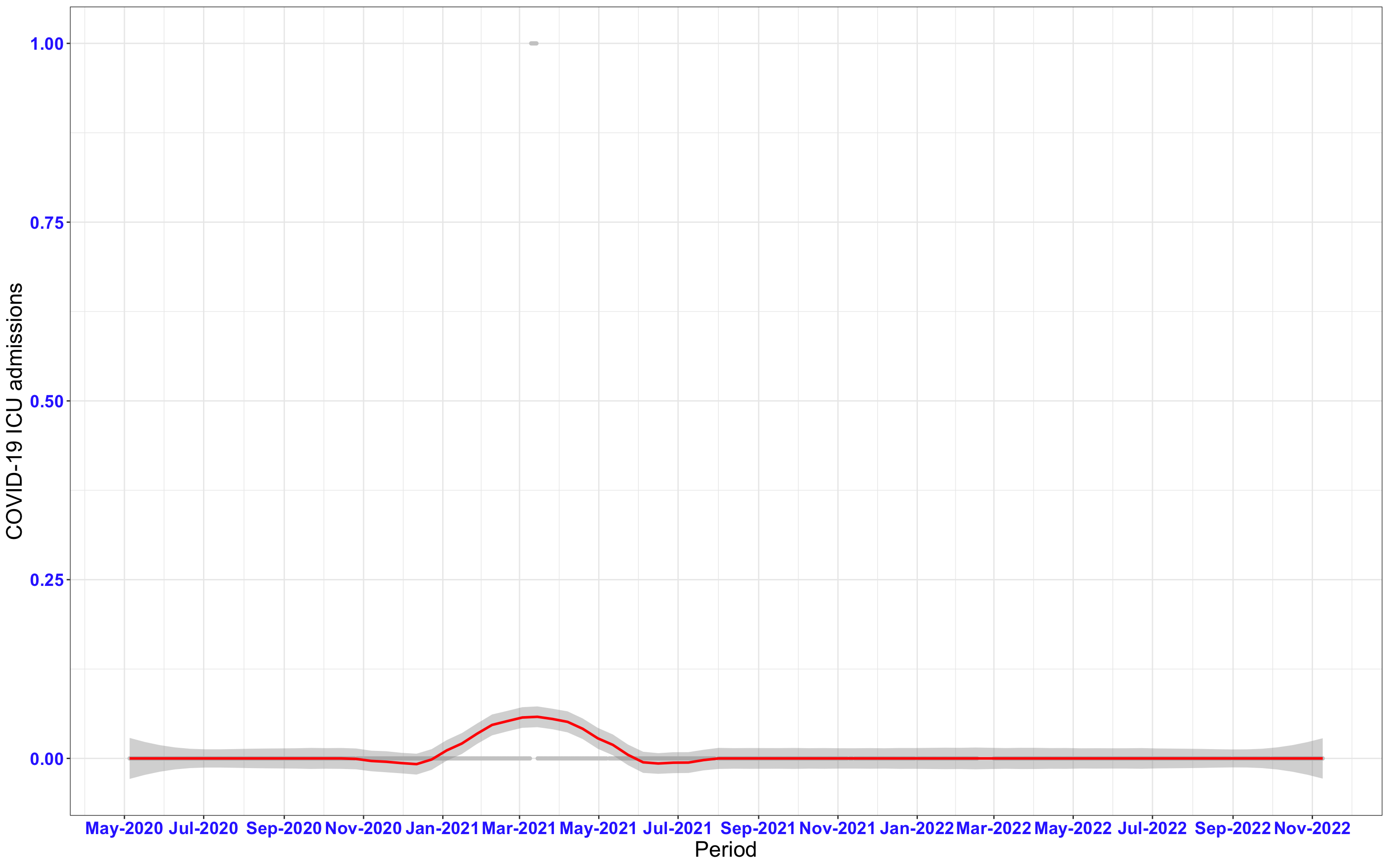
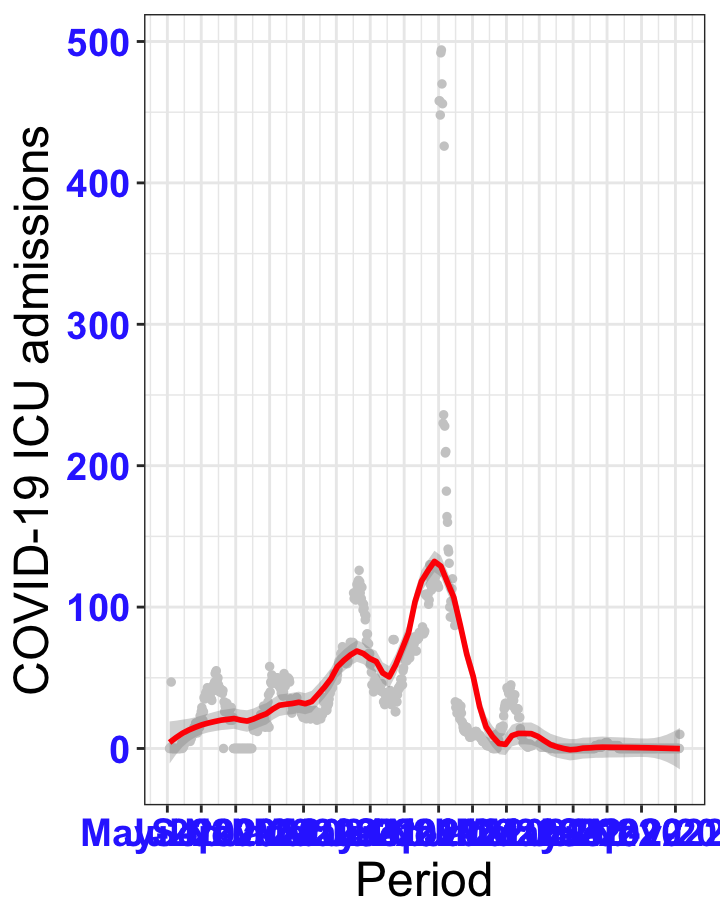
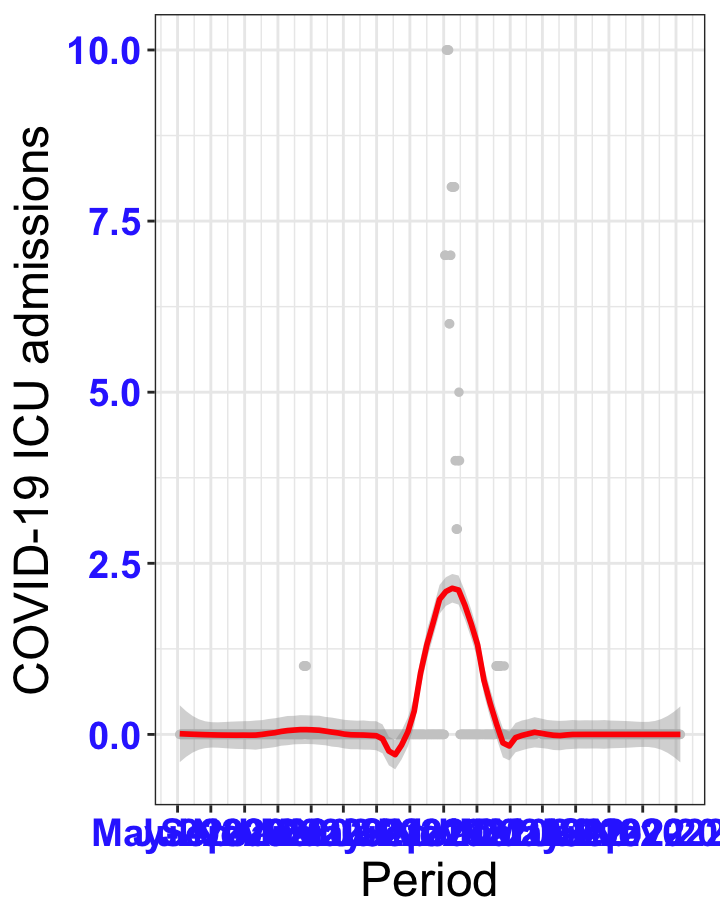
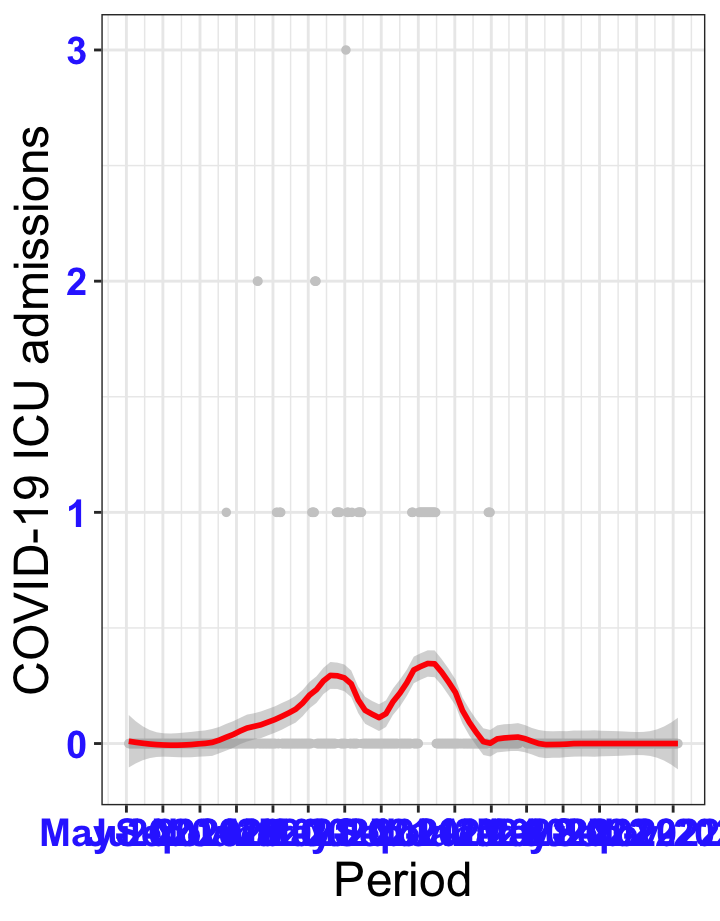
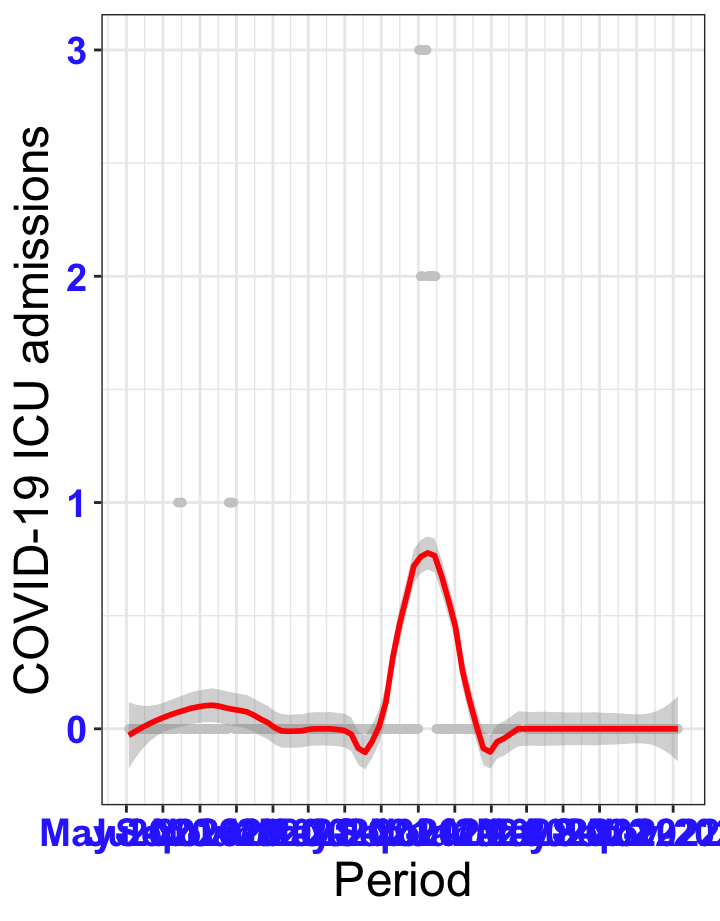
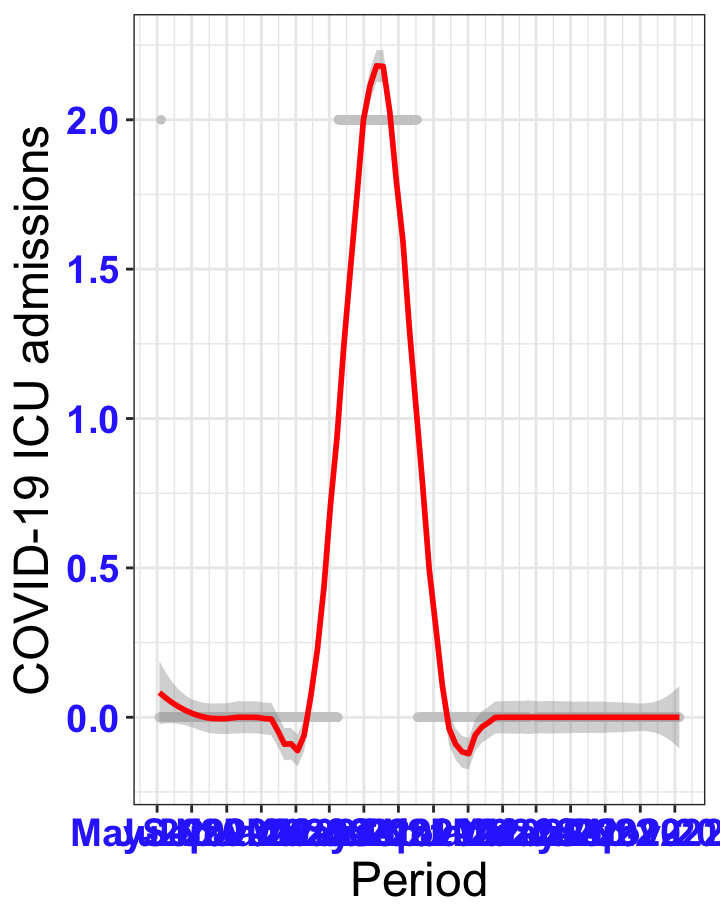
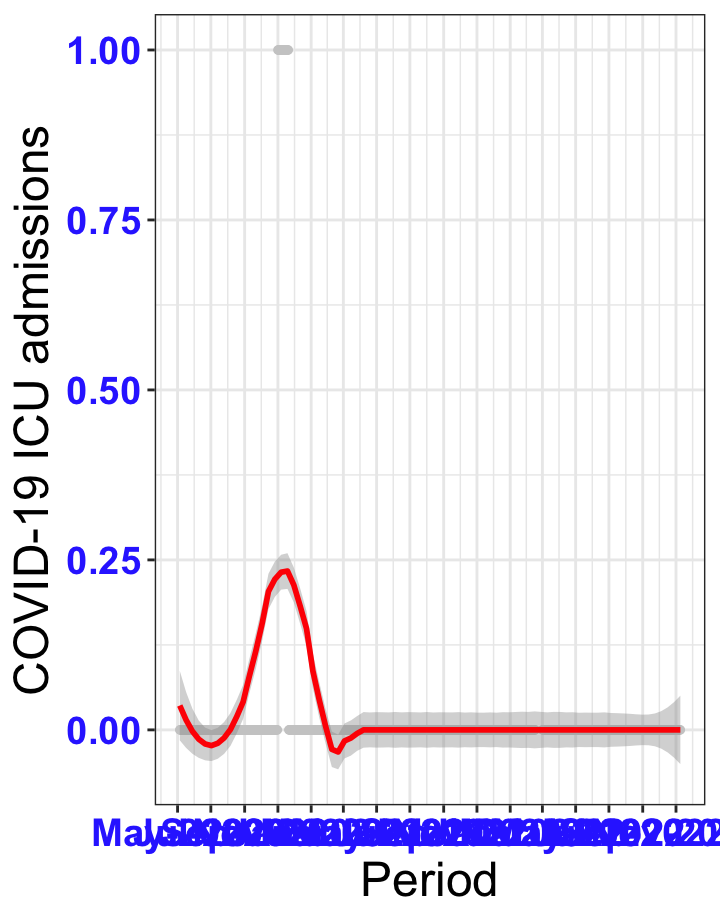
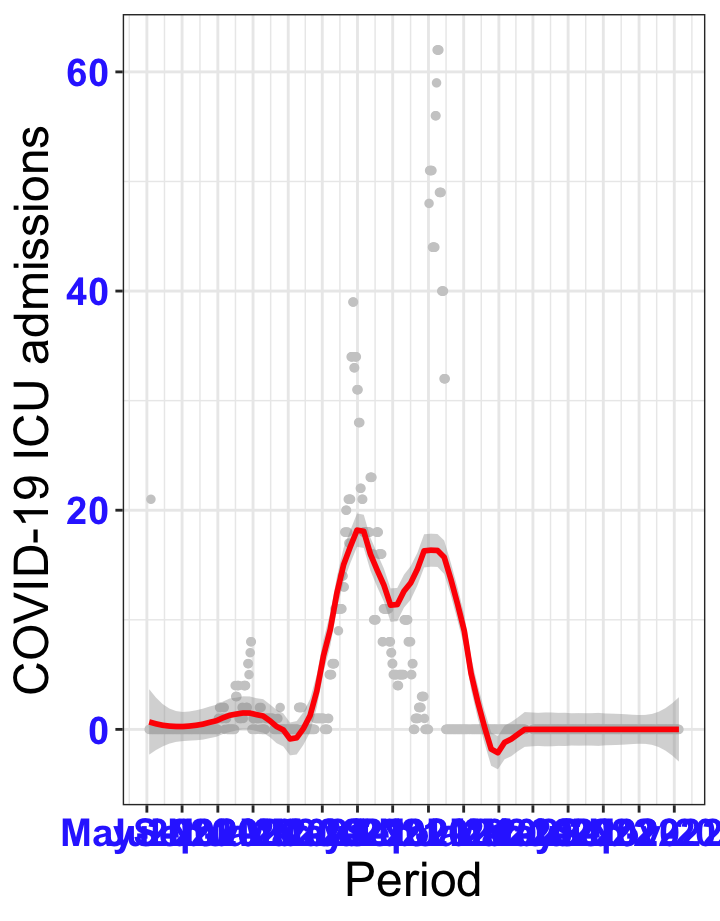
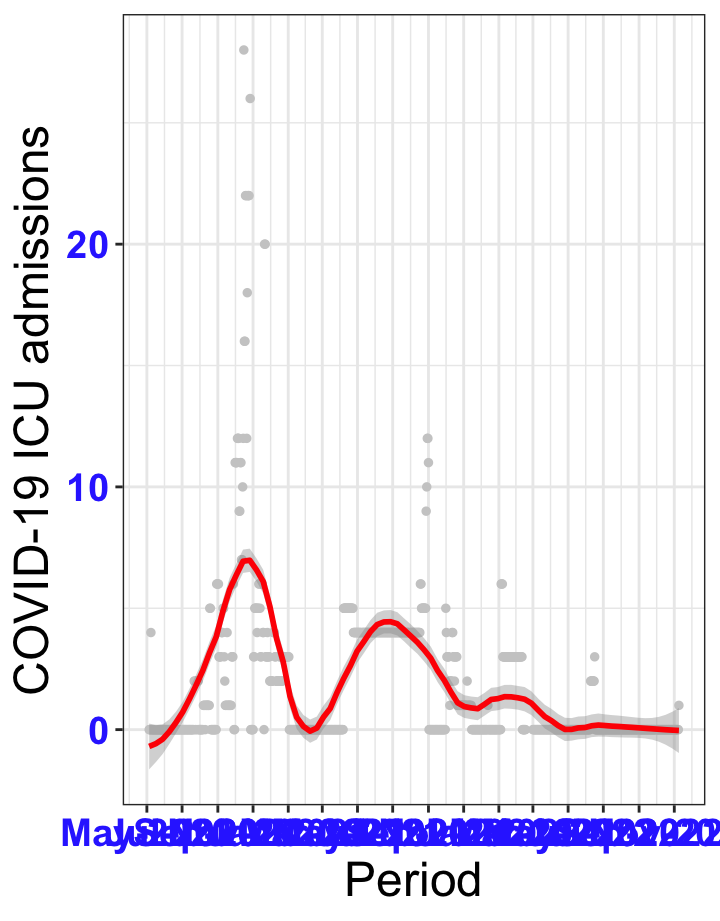
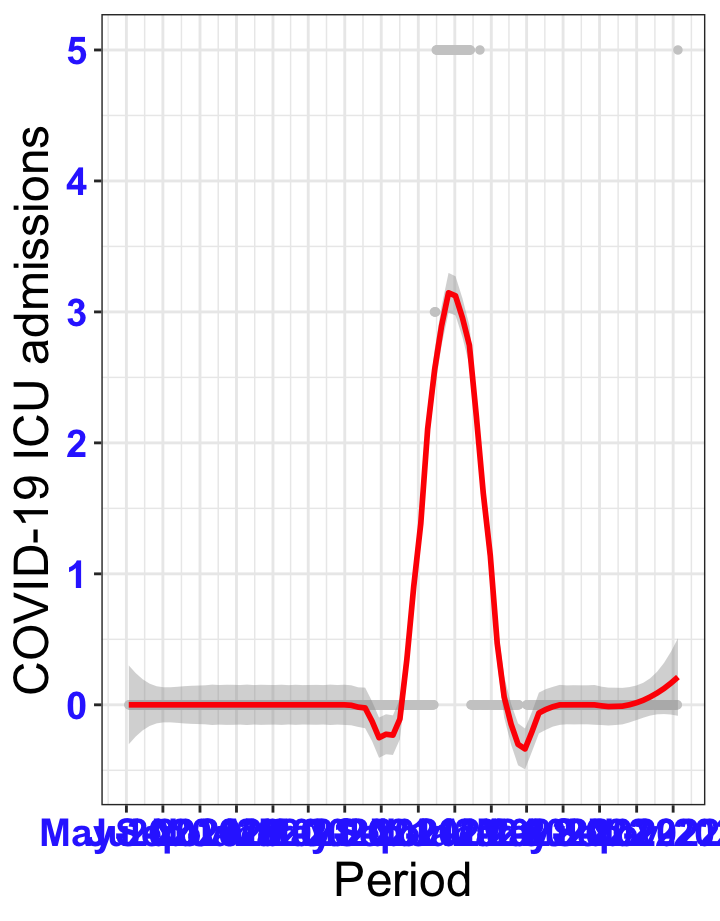
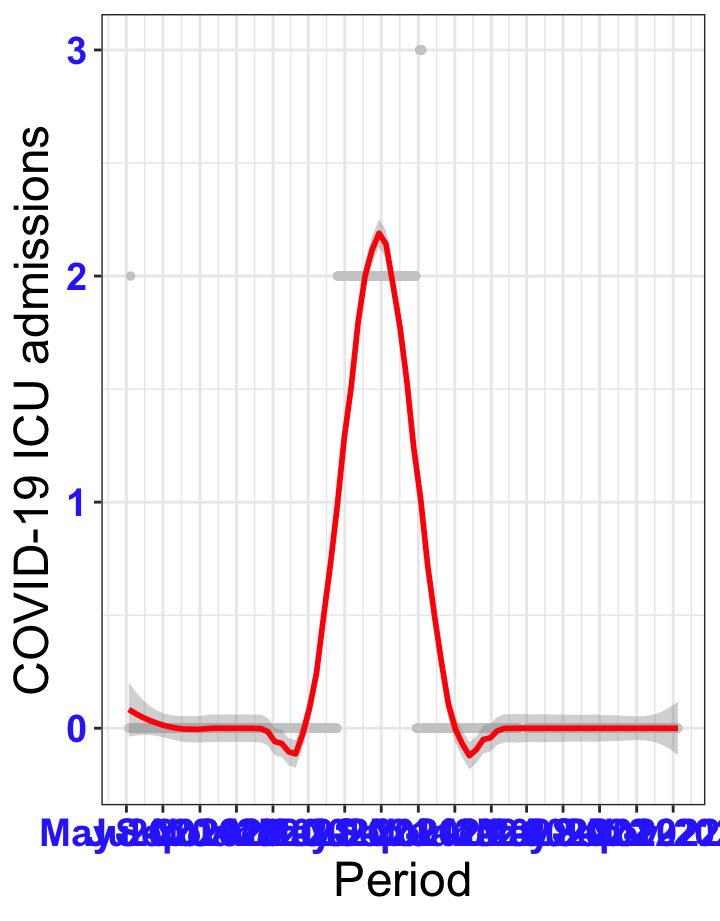
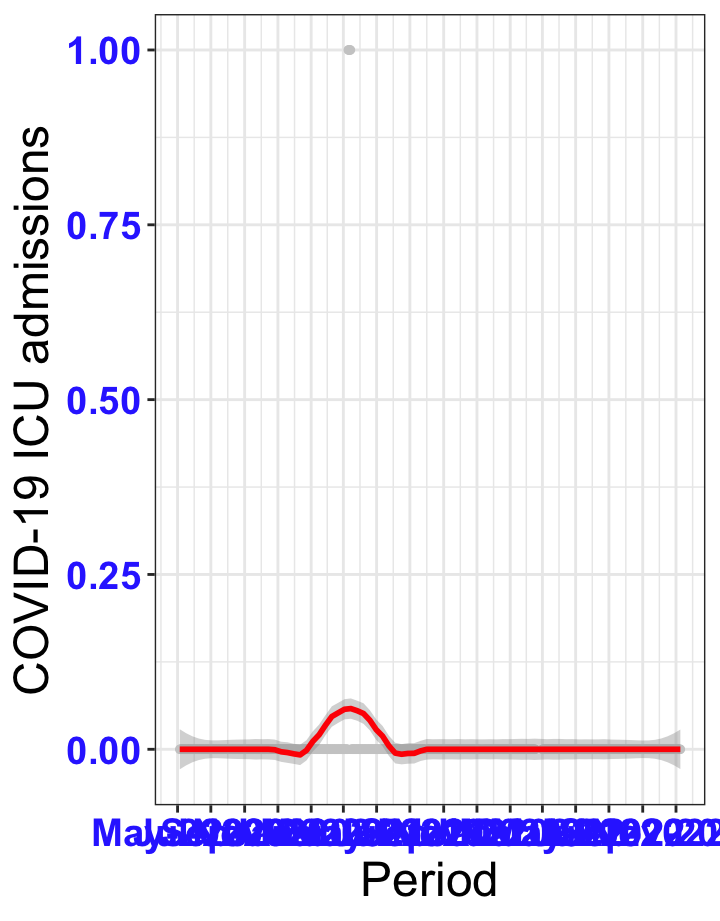
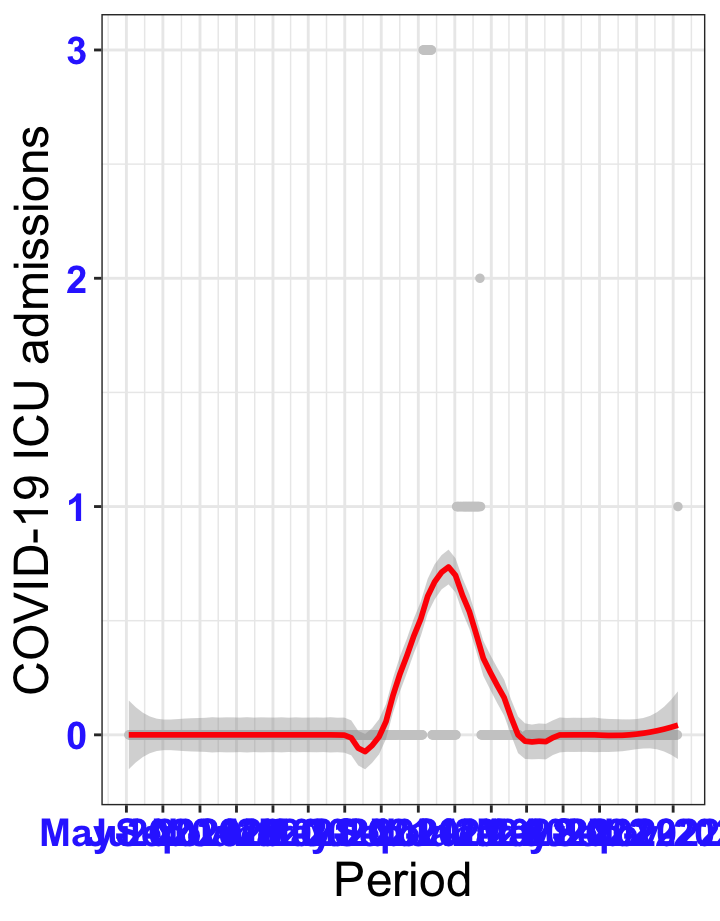
# `r x[1]` {data-navmenu="ICU admissions"}
```{r,fig.width=16, fig.height=10}
library(pander)
graphs1<-list()
for (i in 1:47) {
# Inserts Month titles
#pandoc.header(paste("{data-navmenu="Hospital admissions"}",x[1]), level = 1, "setex")
# Section contents
px<-eval(substitute(ggplot(data_county[data_county$date>as.Date("2020-05-04") & data_county$county%in%x[i],], aes(x=date,y=icu))+geom_point(color="grey80", size=1)+theme_bw()+geom_smooth(span=0.3, colour="red", oob="squish")+scale_x_date(date_labels ="%b-%Y" ,date_breaks = "2 months")+labs(x="Period", y="COVID-19 ICU admissions")+theme(text=element_text(size=18 ), axis.text = element_text(color = "#3339FF", face = "bold")),list(i = i)))
graphs1[[i]]<-px
# adding also empty lines, to be sure that this is valid Markdown
# pander::pandoc.p('')
# pander::pandoc.p('')
}
graphs1[[1]]
```
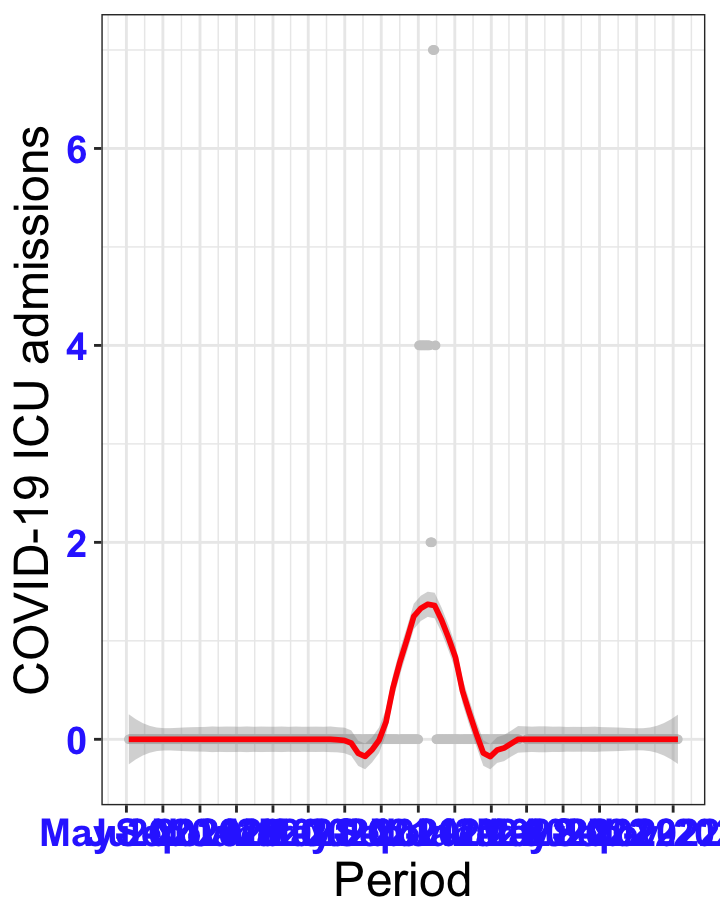
# `r x[2]` {data-navmenu="ICU admissions"}
```{r,fig.width=16, fig.height=10}
graphs1[[2]]
```
# `r x[3]` {data-navmenu="ICU admissions"}
```{r,fig.width=16, fig.height=10}
graphs1[[3]]
```
# `r x[4]` {data-navmenu="ICU admissions"}
```{r,fig.width=16, fig.height=10}
graphs1[[4]]
```
# `r x[5]` {data-navmenu="ICU admissions"}
```{r,fig.width=16, fig.height=10}
graphs1[[5]]
```
# `r x[6]` {data-navmenu="ICU admissions"}
```{r,fig.width=16, fig.height=10}
graphs1[[6]]
```
# `r x[7]` {data-navmenu="ICU admissions"}
```{r,fig.width=16, fig.height=10}
graphs1[[7]]
```
# `r x[8]` {data-navmenu="ICU admissions"}
```{r,fig.width=16, fig.height=10}
graphs1[[8]]
```
# `r x[9]` {data-navmenu="ICU admissions"}
```{r,fig.width=16, fig.height=10}
graphs1[[9]]
```
# `r x[10]` {data-navmenu="ICU admissions"}
```{r,fig.width=16, fig.height=10}
graphs1[[10]]
```
# `r x[11]` {data-navmenu="ICU admissions"}
```{r,fig.width=16, fig.height=10}
graphs1[[11]]
```
# `r x[12]` {data-navmenu="ICU admissions"}
```{r,fig.width=16, fig.height=10}
graphs1[[12]]
```
# `r x[13]` {data-navmenu="ICU admissions"}
```{r,fig.width=16, fig.height=10}
graphs1[[13]]
```
# `r x[14]` {data-navmenu="ICU admissions"}
```{r,fig.width=16, fig.height=10}
graphs1[[14]]
```
# `r x[15]` {data-navmenu="ICU admissions"}
```{r,fig.width=16, fig.height=10}
graphs1[[15]]
```
# `r x[16]` {data-navmenu="ICU admissions"}
```{r,fig.width=16, fig.height=10}
graphs1[[16]]
```
# `r x[17]` {data-navmenu="ICU admissions"}
```{r,fig.width=16, fig.height=10}
graphs1[[17]]
```
# `r x[18]` {data-navmenu="ICU admissions"}
```{r,fig.width=16, fig.height=10}
graphs1[[18]]
```
# `r x[19]` {data-navmenu="ICU admissions"}
```{r,fig.width=16, fig.height=10}
graphs1[[19]]
```
# `r x[20]` {data-navmenu="ICU admissions"}
```{r,fig.width=16, fig.height=10}
graphs1[[20]]
```
# `r x[21]` {data-navmenu="ICU admissions"}
```{r,fig.width=16, fig.height=10}
graphs1[[21]]
```
# `r x[22]` {data-navmenu="ICU admissions"}
```{r,fig.width=16, fig.height=10}
graphs1[[22]]
```
# `r x[23]` {data-navmenu="ICU admissions"}
```{r,fig.width=16, fig.height=10}
graphs1[[23]]
```
# `r x[24]` {data-navmenu="ICU admissions"}
```{r,fig.width=16, fig.height=10}
graphs1[[24]]
```
# `r x[25]` {data-navmenu="ICU admissions"}
```{r,fig.width=16, fig.height=10}
graphs1[[25]]
```
# `r x[26]` {data-navmenu="ICU admissions"}
```{r,fig.width=16, fig.height=10}
graphs1[[26]]
```
# `r x[27]` {data-navmenu="ICU admissions"}
```{r,fig.width=16, fig.height=10}
graphs1[[27]]
```
# `r x[28]` {data-navmenu="ICU admissions"}
```{r,fig.width=16, fig.height=10}
graphs1[[28]]
```
# `r x[29]` {data-navmenu="ICU admissions"}
```{r,fig.width=16, fig.height=10}
graphs1[[29]]
```
# `r x[30]` {data-navmenu="ICU admissions"}
```{r,fig.width=16, fig.height=10}
graphs1[[30]]
```
# `r x[31]` {data-navmenu="ICU admissions"}
```{r,fig.width=16, fig.height=10}
graphs1[[31]]
```
# `r x[32]` {data-navmenu="ICU admissions"}
```{r,fig.width=16, fig.height=10}
graphs1[[32]]
```
# `r x[33]` {data-navmenu="ICU admissions"}
```{r,fig.width=16, fig.height=10}
graphs1[[33]]
```
# `r x[34]` {data-navmenu="ICU admissions"}
```{r,fig.width=16, fig.height=10}
graphs1[[34]]
```
# `r x[35]` {data-navmenu="ICU admissions"}
```{r,fig.width=16, fig.height=10}
graphs1[[35]]
```
# `r x[36]` {data-navmenu="ICU admissions"}
```{r,fig.width=16, fig.height=10}
graphs1[[36]]
```
# `r x[37]` {data-navmenu="ICU admissions"}
```{r,fig.width=16, fig.height=10}
graphs1[[37]]
```
# `r x[38]` {data-navmenu="ICU admissions"}
```{r,fig.width=16, fig.height=10}
graphs1[[38]]
```
# `r x[39]` {data-navmenu="ICU admissions"}
```{r,fig.width=16, fig.height=10}
graphs1[[39]]
```
# `r x[40]` {data-navmenu="ICU admissions"}
```{r,fig.width=16, fig.height=10}
graphs1[[40]]
```
# `r x[41]` {data-navmenu="ICU admissions"}
```{r,fig.width=16, fig.height=10}
graphs1[[41]]
```
# `r x[42]` {data-navmenu="ICU admissions"}
```{r,fig.width=16, fig.height=10}
graphs1[[42]]
```
# `r x[43]` {data-navmenu="ICU admissions"}
```{r,fig.width=16, fig.height=10}
graphs1[[43]]
```
# `r x[44]` {data-navmenu="ICU admissions"}
```{r,fig.width=16, fig.height=10}
graphs1[[44]]
```
# `r x[45]` {data-navmenu="ICU admissions"}
```{r,fig.width=16, fig.height=10}
graphs1[[45]]
```
# `r x[46]` {data-navmenu="ICU admissions"}
```{r,fig.width=16, fig.height=10}
graphs1[[46]]
```
# `r x[47]` {data-navmenu="ICU admissions"}
```{r,fig.width=16, fig.height=10}
graphs1[[47]]
```
```{r, attack rate, eval=FALSE,message=F, warning=F}
linelist<- read_csv("linelist_20221115.csv")%>%
janitor::clean_names()%>%
dplyr::mutate(county_of_residence=str_to_title(county_of_residence))%>%
mutate(county_of_residence=trimws(county_of_residence))%>%
mutate(county_of_residence=recode(county_of_residence, "Egeyo Marakwet"="Elgeyo Marakwet", "Garrissa"="Garissa", "Hoam Bay"="Homa Bay",
"Homabay"="Homa Bay", "Kabete"="Kiambu", "Kitengela"="Kajiado", "Kittui"="Kitui", "Malaba"="Busia", "Murang`A"="Murang'a", "F"="Nairobi", "N"="Nairobi","Kenyan"="Nairobi","Muranga"="Murang'a"))%>%
dplyr::mutate(county_of_residence=ifelse(county_of_residence==c(" ","Kenyan"), "Nairobi", county_of_residence))%>%
filter(county_of_residence!="")%>%
filter(county_of_residence!="/Unknown")%>%
dplyr::mutate(county_of_residence=ifelse(county_of_residence%in%c("American","British","Chinese","Dutch","Eritrean","Filipino","Finnish","German","Indian","Italian","Japanese","South Sudanese","Sri Lankan","Sudanese","Swedish","Ugandan","Zimbabwean","Sweden","Ethiopian"),NA,county_of_residence))
library(lubridate)
linelist$date_of_lab_confirmation1 <- as.Date(linelist$date_of_lab_confirmation,format="%m/%d/%y")
#linelist$date_of_lab_confirmation1 <- excel_numeric_to_date(linelist$date_of_lab_confirmation)
#linelist$date_of_lab_confirmation1 <- convert_to_date(linelist$date_of_lab_confirmation)
linelist1<- linelist%>%
group_by(date_of_lab_confirmation1, county_of_residence)%>%
count()%>%
ungroup()
#linelist <- transform(linelist, date_of_lab_confirmation = as.Date(as.character(date_of_lab_confirmation), "%Y%m%d"))
population<-read_csv("kenya_population.csv")%>%
janitor::clean_names()%>%
filter(county_filter%in%"County")%>%
filter(!ages%in%c("0-4", "10-14", "10-19", "15-19", "20-24", "25-29", "30-34", "35-39", "40-44", "45-49", "5-9", "50-54", "55-59",
"60-64", "65-69", "70-74", "75-79", "80-84", "85-89", "9-14", "90-94", "95-99"))%>%
group_by(county_subcounty)%>%
mutate(total=sum(total, na.rm=T))%>%
ungroup()%>%
select(county_subcounty, total)%>%
distinct()%>%
mutate(county_subcounty=str_to_title(county_subcounty))%>%
mutate(county_subcounty=recode(county_subcounty, "Elgeyo/Marakwet"="Elgeyo Marakwet", "Homa_bay"="Homa Bay", "Nairobi_city"="Nairobi",
"Taita/Taveta"="Taita Taveta", "Tana_river"="Tana River", "Tharaka-Nithi"="Tharaka Nithi", "Trans_nzoia"="Trans Nzoia", "Uasin_gishu"="Uasin Gishu", "West_pokot"="West Pokot"))
xmax1<-as.Date("2021-03-23")-29
xmin1<-as.Date("2021-03-23")+28
linelist1a<-linelist1%>%
#mutate(county_residence1a=recode(county_residence1a,"Muranga"="Murang'a"))%>%
left_join(population, by=c("county_of_residence"="county_subcounty"))%>%
dplyr::filter(date_of_lab_confirmation1>xmax1 & date_of_lab_confirmation1%
# mutate(icu=round(mean(icu)))%>%
# mutate(new_admissions=round(mean(new_admissions)))%>%
# ungroup()%>%
# select(-oxygen, -new_discharges, -admissions, -date)%>%
# distinct()%>%
# mutate(county=recode(county,"Murang'a"="Muranga"))
linelist1b<- linelist1a%>%
group_by(date_of_lab_confirmation1)%>%
mutate(total_county=round(sum(n)/sum(total)*100000))%>%
ungroup()%>%
filter(!is.na(county_of_residence))
linelist1c<- linelist1a%>%
filter(date_of_lab_confirmation1>as.Date("2021-04-05"))%>%
mutate(week=if_else(date_of_lab_confirmation1%
group_by(week, county_of_residence)%>%
mutate(AR1=mean(AR))%>%
ungroup()%>%
select(county_of_residence, week, AR1)%>%
distinct()%>%
pivot_wider(id_cols=county_of_residence, names_from=week, values_from=AR1)
linelist1c$AR2<- linelist1c$`2021-04-20`-linelist1c$`2021-04-12`
linelistd<- linelist1c%>%
filter(AR2>0)
pp<-ggplot()+geom_smooth(data=linelist1b[linelist1b$county_of_residence!="Nairobi",], aes(x=date_of_lab_confirmation1, y=AR, group=county_of_residence), color="red", show.legend = F)+geom_smooth(data=linelist1b, aes(x=date_of_lab_confirmation1, y=total_county) , color="black")+theme_bw()+facet_wrap(~county_of_residence)+geom_vline(xintercept=as.Date("2021-03-26"), linetype=2)+labs(x="Date", y="Attack rate")
pp
```
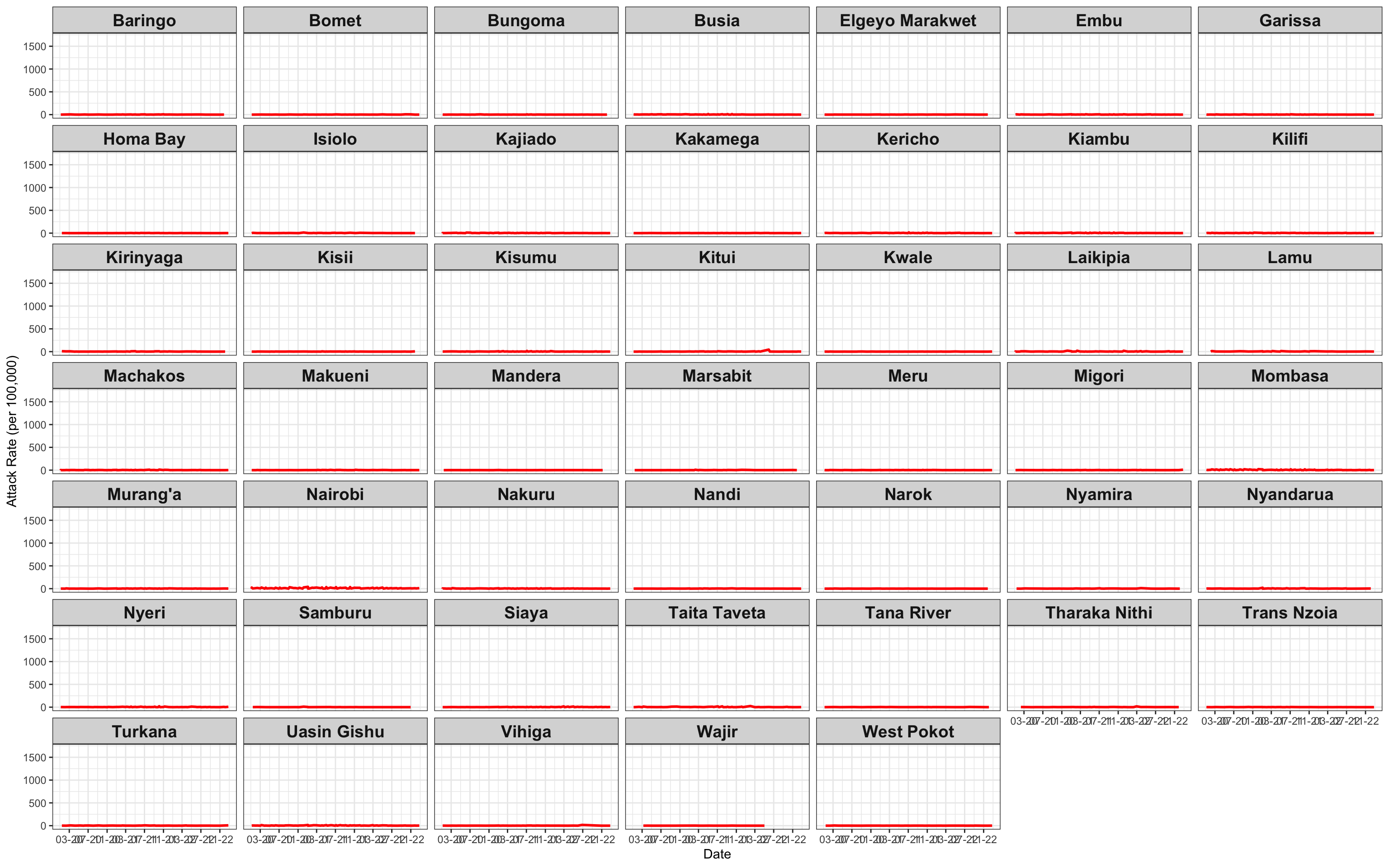
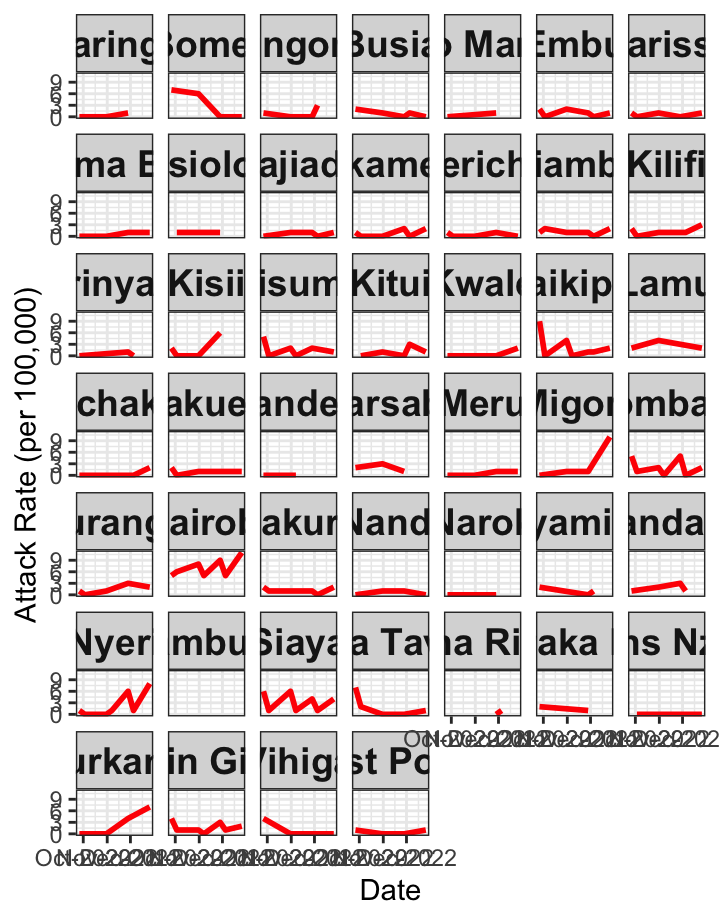
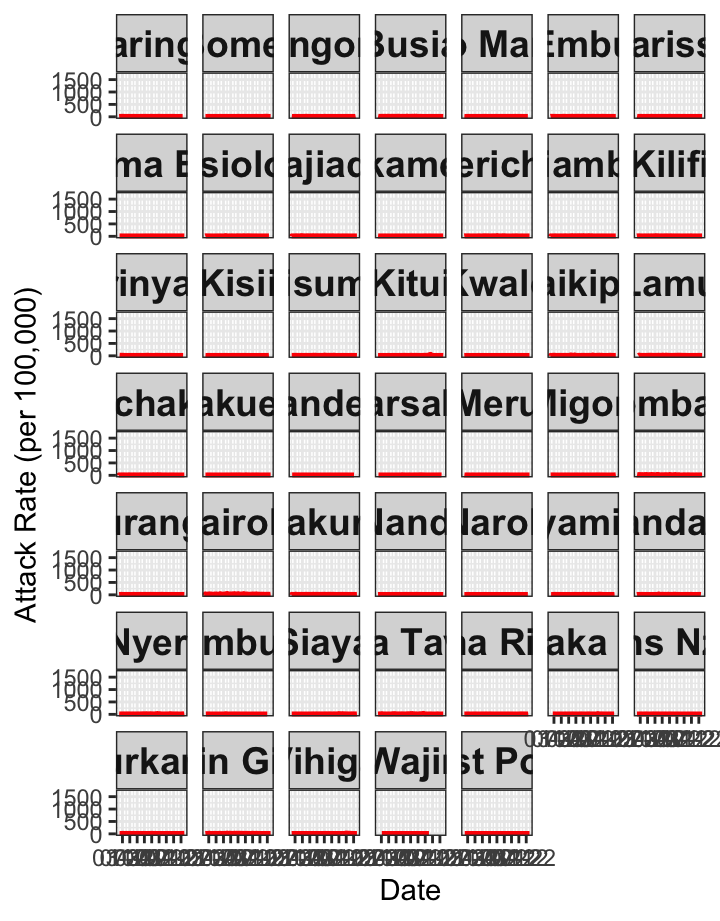
# Attack Rate {data-navmenu="Attack Rate"}
```{r, fig.width=16, fig.height=10}
linelist<- read_csv("linelist_20221115.csv")%>%
janitor::clean_names()%>%
dplyr::mutate(county_of_residence=str_to_title(county_of_residence))%>%
mutate(county_of_residence=trimws(county_of_residence))%>%
mutate(county_of_residence=recode(county_of_residence, "Egeyo Marakwet"="Elgeyo Marakwet", "Garrissa"="Garissa", "Hoam Bay"="Homa Bay",
"Homabay"="Homa Bay", "Kabete"="Kiambu", "Kitengela"="Kajiado", "Kittui"="Kitui", "Malaba"="Busia", "Murang`A"="Murang'a", "F"="Nairobi", "Burundi"="Nairobi", "N"="Nairobi","Kenyan"="Nairobi","Muranga"="Murang'a"))%>%
dplyr::mutate(county_of_residence=ifelse(county_of_residence=="", "Nairobi", county_of_residence))%>%
filter(county_of_residence!="")%>%
mutate(date_of_lab_confirmation1=as.Date(date_of_lab_confirmation, "%d/%m/%Y"))%>%
filter(county_of_residence!="/Unknown")%>%
dplyr::mutate(county_of_residence=ifelse(county_of_residence%in%c("American","British","Chinese","Dutch","Eritrean","Filipino","Finnish","German","Indian","Italian","Japanese","South Sudanese","Sri Lankan","Sudanese","Swedish","Ugandan","Zimbabwean","Ethiopian","Sweden","Usa"),NA,county_of_residence))
linelist$date_of_lab_confirmation1 <- as.Date(linelist$date_of_lab_confirmation,format="%d/%m/%y")
#linelist$date_of_lab_confirmation1 <- janitor::excel_numeric_to_date( linelist$date_of_lab_confirmation)
# linelist11<- linelist%>%
# group_by(date_of_lab_confirmation1, county_of_residence, outcome_death_discharge_still_in_hospital)%>%
# count()%>%
# ungroup()%>%
# filter(date_of_lab_confirmation1>as.Date("2021-05-10"))%>%
# mutate(outcome_death_discharge_still_in_hospital=str_to_sentence(outcome_death_discharge_still_in_hospital))%>%
# filter(outcome_death_discharge_still_in_hospital%in%"Dead")%>%
# pivot_wider(id_cols=county_of_residence, names_from = date_of_lab_confirmation1,values_from = n)
#
# write_csv(linelist11, "covid_deaths_County_20210411.csv")
linelist1<- linelist%>%
mutate(week=epiweek(date_of_lab_confirmation1))%>%
mutate(year=str_sub(date_of_lab_confirmation1, 1, 4))%>%
group_by(week, year, county_of_residence)%>%
count()%>%
ungroup()
#linelist <- transform(linelist, date_of_lab_confirmation = as.Date(as.character(date_of_lab_confirmation), "%Y%m%d"))
population<-read_csv("kenya_population.csv")%>%
janitor::clean_names()%>%
filter(county_filter%in%"County")%>%
filter(!ages%in%c("0-4", "10-14", "10-19", "15-19", "20-24", "25-29", "30-34", "35-39", "40-44", "45-49", "5-9", "50-54", "55-59",
"60-64", "65-69", "70-74", "75-79", "80-84", "85-89", "9-14", "90-94", "95-99"))%>%
group_by(county_subcounty)%>%
mutate(total=sum(total, na.rm=T))%>%
ungroup()%>%
select(county_subcounty, total)%>%
distinct()%>%
mutate(county_subcounty=str_to_title(county_subcounty))%>%
mutate(county_subcounty=recode(county_subcounty, "Elgeyo/Marakwet"="Elgeyo Marakwet", "Homa_bay"="Homa Bay", "Nairobi_city"="Nairobi",
"Taita/Taveta"="Taita Taveta", "Tana_river"="Tana River", "Tharaka-Nithi"="Tharaka Nithi", "Trans_nzoia"="Trans Nzoia", "Uasin_gishu"="Uasin Gishu", "West_pokot"="West Pokot"))
linelist1a<-linelist1%>%
#mutate(county_residence1a=recode(county_residence1a,"Muranga"="Murang'a"))%>%
left_join(population, by=c("county_of_residence"="county_subcounty"))#%>%
#dplyr::filter(date_of_lab_confirmation1>xmax1 & date_of_lab_confirmation1%
# mutate(icu=round(mean(icu)))%>%
# mutate(new_admissions=round(mean(new_admissions)))%>%
# ungroup()%>%
# select(-oxygen, -new_discharges, -admissions, -date)%>%
# distinct()%>%
# mutate(county=recode(county,"Murang'a"="Muranga"))
# linelist1b<- linelist1a%>%
# filter(date>"2021-07-01")%>%
# group_by(week1, year)%>%
# mutate(total_county=round(sum(n)/sum(total)*100000))%>%
# ungroup()
# linelist1c<- linelist1a%>%
# filter(date_of_lab_confirmation1>as.Date("2021-04-05"))%>%
# mutate(week=if_else(date_of_lab_confirmation1%
# group_by(week, county_of_residence)%>%
# mutate(AR1=mean(AR))%>%
# ungroup()%>%
# select(county_of_residence, week, AR1)%>%
# distinct()%>%
# pivot_wider(id_cols=county_of_residence, names_from=week, values_from=AR1)
#
# linelist1c$AR2<- linelist1c$`2021-04-20`-linelist1c$`2021-04-12`
#
# linelistd<- linelist1c%>%
# filter(AR2>0)
#kableExtra::kable(linelistd%>%dplyr::select(county_of_residence,AR2), col.names = c("County", "Attack Rate"), caption="Counties with an increasing attack rate over the past one week" )
linelist1a<- linelist1a%>%
dplyr::mutate(year=as.numeric(year))%>%
dplyr::mutate(week1=ifelse(year==2020, week-1, week))%>%
dplyr::mutate(week1= ifelse(week1==53, 1, week1))%>%
filter(!is.na(county_of_residence))
linelist1a$date<- as.Date(paste(linelist1a$year, linelist1a$week1, 1, sep="-"), "%Y-%U-%u")
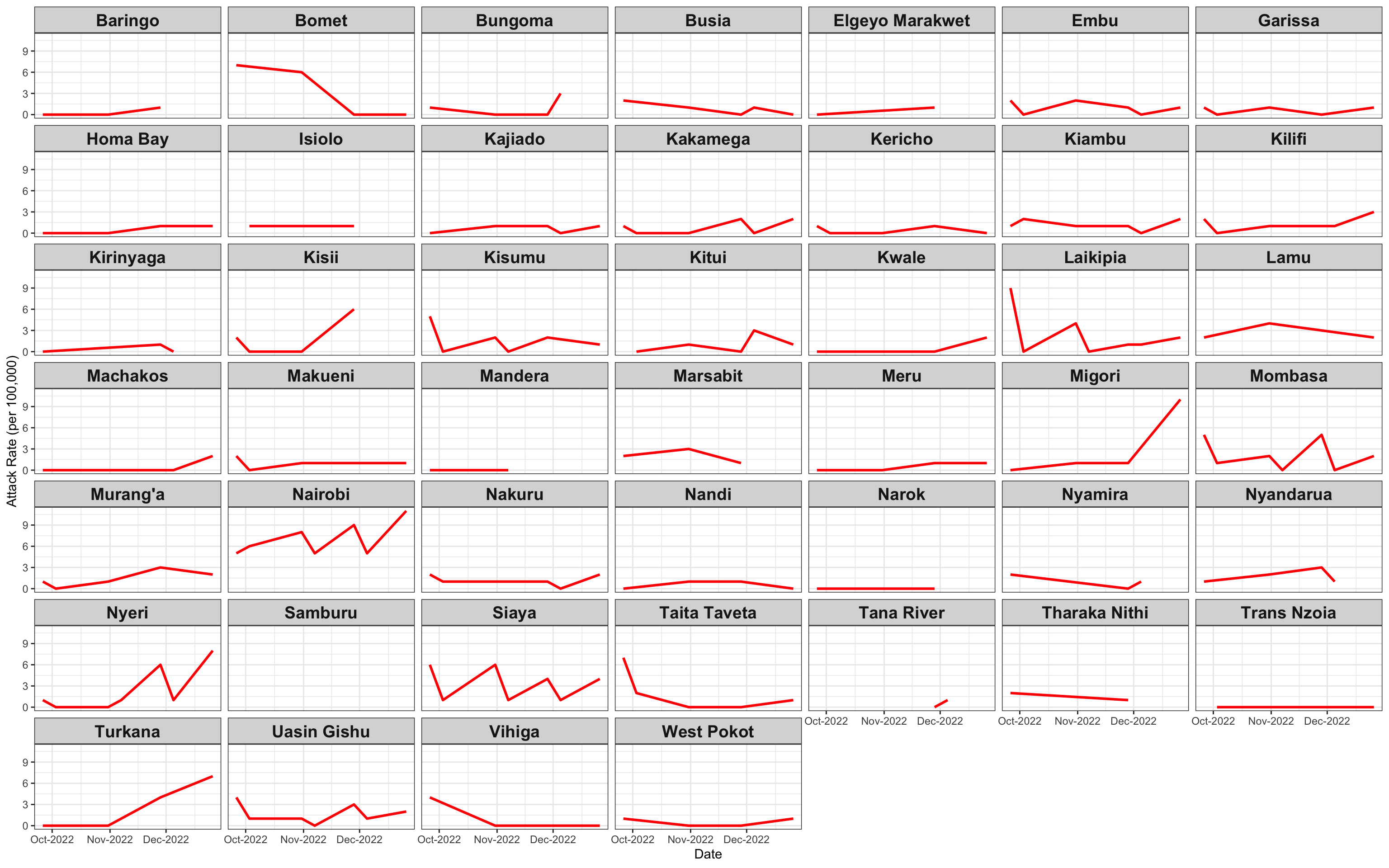
ggplot(linelist1a, aes(x=date, y=AR))+facet_wrap(~county_of_residence)+geom_line(color="red", size=1)+theme_bw()+labs(x="Date", y="Attack Rate (per 100,000)")+
theme(strip.text.x = element_text(size = 14, face="bold"))+scale_x_date(date_breaks="4 months", date_labels="%m-%y")
# ggplot(linelist1a, aes(x=date, y=n))+facet_wrap(~county_of_residence, scales ="free_y")+geom_line(color="red", size=1)+theme_bw()+labs(x="Date", y="Number of cases per week")+
# theme(strip.text.x = element_text(size = 14, face="bold"))+scale_x_date(date_breaks="6 months", date_labels="%b-%Y")
#
# ggsave("county_cases1.png", width=16, height=8)
```
# Attack Rate (Last 8 weeks) {data-navmenu="Attack Rate"}
```{r,fig.width=16, fig.height=10}
last8<-Sys.Date()-57
linelist1a_x <- linelist1a%>%filter(date>last8)
ggplot(linelist1a_x, aes(x=date, y=AR))+facet_wrap(~county_of_residence)+geom_line(color="red", size=1)+theme_bw()+labs(x="Date", y="Attack Rate (per 100,000)")+theme(strip.text.x = element_text(size = 14, face="bold"))+scale_x_date( date_labels="%b-%Y")
```
```{r, eval=F, include=F}
## forecasting models
kisumu<- data%>%
filter(county%in%"Kisumu")%>%
select(county, date, admissions)%>%
distinct()
library(fpp3)
kisumu_df<- kisumu%>%
mutate(date= yearmonth(as.character(date))) %>%
group_by(date)%>%
mutate(admissions=sum(admissions, na.rm=T))%>%
ungroup()%>%
distinct()%>%
as_tsibble()%>%
mutate(covid="Yes")
kisumu_df1<- kisumu%>%
as_tsibble()%>%
mutate(covid="Yes")
fun_reconcile<- function(R, M, B, BU="bu", OLS="ols", MINT="mint"){
LHS= "admissions"
RHS= R
model_spec= as.formula(paste0(LHS, RHS, sep=""))
df_hts %>%
filter( Date < yearmonth("2020 May")) %>%
model(base = model_spec %>% M) %>%
reconcile(
bu = bottom_up(base), ols = min_trace(base, method = "ols"), mint = min_trace(base, method = "mint_cov")) %>%
#https://stackoverflow.com/questions/35023375/r-renaming-passed-columns-in-functions
rename({{B}} :=base) %>%
#https://www.tidyverse.org/blog/2020/02/glue-strings-and-tidy-eval/
rename("{{B}}_{BU}":= bu) %>% rename("{{B}}_{OLS}" := ols) %>% rename("{{B}}_{MINT}" := mint)
}
m_ets<- fun_reconcile("~ error() + trend() + season()", ETS, ets)
m_arima<- fun_reconcile("~ pdq() + PDQ()", ARIMA, arima)
m_arima_covid<- fun_reconcile("~ Covid", ARIMA, arima_covid)
m_arima_covidL1<- fun_reconcile("~ Covid +lag(Covid)", ARIMA, arima_covidL1)
m_arima_covidL2<- fun_reconcile("~ Covid +lag(Covid,1)", ARIMA, arima_covidL2)
m_arima_covidL3<- fun_reconcile("~ Covid +lag(Covid,2)", ARIMA, arima_covidL3)
# save models
save(m_ets, m_arima, m_arima_covid, m_arima_covidL1, m_arima_covidL2, m_arima_covidL3, file = "3bClassic")
kisumu_df1a<-tsibble::fill_gaps(kisumu_df1)
fit <- kisumu_df1a %>%
model(
ets = ETS(admissions~ trend("A")),
arima = ARIMA(admissions)
)
fit %>%
coef()
fit %>%
accuracy() %>%
arrange(MASE)
fc <- fit %>%
dplyr::select(arima)%>%
forecast(h = "30 days")
fc %>%
hilo(level = c(80, 95))
fc %>%
autoplot(kisumu_df1, size=1)+theme_bw()+labs(x="Period", y="Per capita monthly water consumption")+
theme(text=element_text(size=14, face="bold"))
library(astsa)
library(forecast)
library(dplyr)
library(zoo)
kisumu_ts <- ts(kisumu_df[,3], frequency=12, start=c(2020,3))
# To forecast the counterfactual, model data excluding post-intervention time period
model_kisumu <- Arima(window(kisumu_ts, end=c(2021,6)), order=c(2,1,0))
# Forecast 12 months post-intervention and convert to time series object
fc_kisumu <- forecast(model_kisumu, h=12, level=95)
fc_kisumu1<- data.frame(fc_kisumu)
fc_kisumu_ts <- ts(as.numeric(fc_kisumu$mean), start=c(2021,7), frequency=12)
kisumu_ts2 <- ts.union(kisumu_ts, fc_kisumu_ts)
kisumu_ts2a<- data.frame(kisumu_ts2)
kisumu_ts2a$year<- c(rep(c(2020), 10), 2021,2021, 2021,2021,2021, 2021, 2021,2021, 2021,2021,2021, 2021,2022,2022, 2022,2022,2022, 2022)
kisumu_ts2a$date<- seq.Date(as.Date("2020-03-01"), as.Date("2022-06-01"), by="1 month")
kisumu_ts2b<- kisumu_ts2a%>%
select(date, kisumu_ts, fc_kisumu_ts)
kisumu_ts2b1<- reshape2::melt(kisumu_ts2b, id=1)%>%
mutate(variable=recode(variable, "kisumu_ts"="Observed values", "fc_kisumu_ts"="Predicted values"))%>%
filter(!is.na(value))
fc_kisumu1<- fc_kisumu1%>%
mutate(date=seq.Date(as.Date("2021-07-01"), as.Date("2022-06-01"), by="1 month"))
kisumu1<-ggplot()+geom_line(data=kisumu_ts2b1, aes(x=date, y=value), color="#4575b4", size=1)+geom_point(data=kisumu_ts2b1, aes(x=date, y=value), color="#4575b4", size=1)+geom_ribbon(data=fc_kisumu1, aes(x=date, ymin = Lo.95, ymax = Hi.95), fill = "grey70", alpha=0.4)+geom_line(data=fc_kisumu1, aes(x=date, y=Point.Forecast),color="#d73027", size=1)+geom_point(data=fc_kisumu1, aes(x=date, y=Point.Forecast),color="#d73027", size=1)+theme_bw()+labs(x="Period", y="Number of admissions", color="", title="Number of COVID-19 admissions in Kisumu County")+scale_y_continuous(label=scales::comma)+scale_x_date(date_breaks = "2 months", date_labels = "%b-%Y")+xlim(as.Date("2020-03-01"), as.Date("2021-08-01"))
deaths<- linelist%>%
mutate(outcome=ifelse(outcome_death_discharge_still_in_hospital%in%c("Dead", "DEAD"), "Dead", "Alive"))%>%
select(county_of_residence, date_of_lab_confirmation1, outcome)%>%
group_by(county_of_residence,date_of_lab_confirmation1, outcome)%>%
count()%>%
ungroup()%>%
filter(outcome%in%"Dead")
ggplot(deaths, aes(x=date_of_lab_confirmation1, y=n))+facet_wrap(~county_of_residence)+geom_line()+scale_x_date(date_breaks = "4 months", date_labels = "%b-%y")+theme_bw()+labs(x="Period(month-year)", y="Frequency")
ggsave("covid_deaths.png", width=14, height=7)
write_csv(deaths, "covid_deaths.csv")
```
# Download {data-navmenu="Download"}
```{r}
library(DT)
df_daily <- coronavirus %>%
dplyr::filter(date >= "2020-03-10") %>%
dplyr::filter(country == "Kenya") %>%
dplyr::group_by(date, type) %>%
dplyr::summarise(total = sum(cases, na.rm = TRUE)) %>%
tidyr::pivot_wider(
names_from = type,
values_from = total
) %>%
dplyr::arrange(date) %>%
dplyr::ungroup() %>%
#dplyr::mutate(active = confirmed - death - recovered) %>%
dplyr::mutate(active = confirmed - death) %>%
dplyr::mutate(
confirmed_cum = cumsum(confirmed),
recovered_cum = cumsum(recovery),
death_cum = cumsum(death),
active_cum = cumsum(active)
)
dt1 <- datatable(df_daily, rownames=F,
caption = "COVID-19 numbers for Kenya: Death, recovered, active cases and cumulative cases for each",
# filter = 'top',
callback=JS('$("a.buttons-collection").css("background","#008CBA");
$("a.buttons-collection").css("font-size","15px");
$("a.buttons-collection").css("border-radius", "6px");
$("a.buttons-collection").css("margin-right","0px");
return table;'),
extensions = 'Buttons',
options = list(searching=T,
paging = T,
bInfo = F,
columnDefs = list(list(className = 'dt-center'),
list(className = 'dt-center'),
list(className = 'dt-center'),
list(className = 'dt-center')),
pageLength = 10,
initComplete = JS("function(settings, json) {",
"$(this.api().table().header()).css({'background-color': 'white', 'color': '#003333'});",
"}"),
dom = 'Bfrtip',
buttons = list(
list(extend = 'collection',
buttons = c('excel', 'csv'),
text = 'Download Data')
)
)
)
dt1
```